 |
 |
| Plant Pathol J > Volume 30(2); 2014 > Article |
Abstract
Although multiple transcription factors (TFs) have been characterized via mutagenesis to understand their roles in controlling pathogenicity and infection-related development in Magnaporthe oryzae, the causal agent of rice blast, if and how forkhead-box (FOX) TFs contribute to these processes remain to be characterized. Four putative FOX TF genes were identified in the genome of M. oryzae, and phylogenetic analysis suggested that two of them (MoFKH1 and MoHCM1) correspond to Ascomycota-specific members of the FOX TF family while the others (MoFOX1 and MoFOX2) are Pezizomycotina-specific members. Deletion of MoFKH1 (ΔMofkh1) resulted in reduced mycelial growth and conidial germination, abnormal septation and stress response, and reduced virulence. Similarly, ΔMohcm1 exhibited reduced mycelial growth and conidial germination. Conidia of ΔMofkh1 and ΔMohcm1 were more sensitive to one or both of the cell cycle inhibitors hydroxyurea and benomyl, suggesting their role in cell cycle control. On the other hand, loss of MoFOX1 (ΔMofox1) did not show any noticeable changes in development, pathogenicity, and stress response. Deletion of MoFOX2 was not successful even after repeated attempts. Taken together, these results suggested that MoFKH1 and Mo-HCM1 are important in fungal development and that MoFKH1 is further implicated in pathogenicity and stress response in M. oryzae.
The rice blast fungus Magnaporthe oryzae is a serious threat to rice production worldwide, causing losses enough to feed 60 million people every year (Dean et al., 2005). When typically three-celled conidia of M. oryzae, the primary inoculum, land on the surface of rice, they germinate and form specialized infection structures, such as the appressorium and invasive hyphae, in sequence to invade rice tissue. Following successful in planta proliferation, M. oryzae produces conidia to initiate next cycle (Ebbole, 2007; Talbot, 1995). Several genes required for appressorium formation (Soanes et al., 2012; Talbot, 2003; Wilson and Talbot, 2009) and those expressed in planta (Kim et al., 2010) have been identified. Systematic identification and characterization of candidate genes associated with infection-related traits have been greatly facilitated by the release of annotated genome sequences of M. oryzae (Dean et al., 2005).
Recently, a number of M. oryzae genes encoding transcription factors (TFs) have been characterized to determine their involvement in controlling pathogenicity and development. MoCRZ1, a calcineurin-responsive C2H2-type zinc finger TF, regulates growth and pathogenicity in response to Ca2+-dependent signaling (Choi et al., 2009). The MIG1 protein, a MADS box TF, functions downstream of the MAP kinase signaling pathway and is required for infectious growth (Mehrabi et al., 2008). Homeobox genes MoHOX2, MoHOX7, and MST12 (=MoHOX8) encode developmental stage-specific TFs that control asexual reproduction, appressorium formation, and host penetration, respectively (Kim et al., 2009). The bZIP TF MoAP1 is critical for oxidative stress response, development, and pathogenicity (Guo et al., 2011). Gene expression analysis of most M. oryzae TF genes under diverse growth conditions and stresses and developmental stages provided comprehensive insights into the transcriptional regulatory network controlling and coordinating its growth, development, pathogenicity and stress responses (Park et al., 2013).
In the current study, we report the characterization of three members of the forkhead-box (FOX) TF gene family, which was named after the Drosophila melanogaster gene responsible for two spiked-head mutant (Jürgens et al., 1984; Weigel et al., 1989), in M. oryzae. FOX TFs contain a DNA binding domain of about 110 amino acids, which consists of three α-helices, three β-strands, and two ‘wing’ regions and forms a helix-turn-helix structure called ‘winged helix’. Most FOX TFs bind to DNA as a monomer and are involved in controlling a wide range of biological processes such as cell cycle progression, growth and differentiation (Carlsson and Mahlapuu, 2002). The genes for FOX TFs have been found in the genomes of animals and fungi, but not plants (Shimeld et al., 2010). In Saccharomyces cerevisiae, four FOX TFs, FKH1, FKH2, HCM1, and FHL1, have been characterized. FKH1 and FKH2 are regulators of genes involved in the G2 and M phases of cell cycle, and disruption of FKH1 and FKH2 caused impaired cell division and abnormal morphology (Koranda et al., 2000; Kumar et al., 2000; Zhu et al., 2000). HCM1 takes part in controlling the S phase and chromosome segregation (Pramila et al., 2006), while FHL1 seems to control rRNA processing (Hermann-Le Denmat et al., 1994). The four FOX TF genes in Schizosaccharomyces pombe, including fkh2+, sep1+, mei4+ and fhl1+, also are important for cell cycle control and morphogenesis. Fkh2, a homolog of S. cerevisiae FKH1 and FKH2, regulates the periodic expression of M- and G1-phase genes and is required for normal cell division (Bulmer et al., 2004). The sep1+ gene controls cell separation and is transcribed in cell cycle dependent manner (Ribár et al., 1999). The mei4+ gene encodes a meiosis-specific TF that is transcribed after premeiotic DNA replication (Horie et al., 1998) and fhl1+, a homolog of S. cerevisiae FHL1, shows a non-periodical expression pattern and participates in cell morphogenesis (Szilagyi et al., 2005). CaFKH2 in Candida albicans controls yeast and true-hyphal morphogenesis. Mutant lacking CaFKH2 formed pseudohyphae and lost its virulence (Bensen et al., 2002). In Aspergillus nidulans, the fkhF and fkhE genes encode FOX TFs and are important for asexual development (Park et al., 2010). Lee et al. (2005) reported that A. nidulans fhpA gene is a possible regulator of sexual development.
To reveal the roles of FOX TFs in M. oryzae, we conducted phylogenetic analysis and functionally characterized FOX TF genes in M. oryzae. MoFKH1 deletion (ΔMofkh1) affected mycelial growth, conidial germination, septation, stress response, and pathogenicity. Loss of MoHCM1 (ΔMohcm1) caused defects in mycelial growth and conidial germination. ΔMofkh1 and ΔMohcm1 also showed higher sensitivity to cell cycle inhibitors. However, ΔMofox1 was indistinguishable from the wild-type and MoFOX2 deletion was not successful even after repeated attempts. Overall, our results revealed roles of MoFKH1 and MoHCM1 in fungal development, pathogenicity, and stress response, which will contribute to elucidating the regulatory mechanism of M. oryzae infection.
M. oryzae wild-type strain KJ201 was obtained from the Center for Fungal Genetic Resources (CFGR, http://knrrb.knrrc.or.kr/index.jsp?rrb=cfgr). For conidial production, wild-type and mutant strains were grown at 25°C on oatmeal agar (OMA, 50 g of oatmeal and 25 g of agar per liter) or V8 juice agar (V8A, 80 ml of V8 juice, 310 μl of 10 N NaOH, and 15 g of agar per liter) under continuous fluorescent light. Conidia were harvested by rubbing the culture surface with sterilized distilled water followed by filtration through Miracloth (Calbiochem, USA). DNA was isolated from mycelia grown in liquid complete medium (CM, 6 g of yeast extract, 6 g of Casamino acids, and 10 g of sucrose per liter). To evaluate mycelial growth, modified complete agar (MCA, 10 g of glucose, 2 g of peptone, 1 g of yeast extract, 1 g of Casamino acids, 6 g of NaNO3, 0.5 g of KCl, 0.5 g of MgSO4, 1.5 g of KH2PO4, and 15 g of agar per liter supplemented with trace elements and vitamins) and modified minimal agar (MMA, MCA without peptone, yeast extract, and Casamino acids) (Talbot et al., 1993) were used.
Via InterProScan using the term IPR001766, which corresponds to the DNA binding domain of FOX TFs, putative FOX TFs were identified. Sequences of FOX TFs in various fungi were obtained from the Fungal Transcription Factor Database (FTFD, http://ftfd.snu.ac.kr) (Park et al., 2008) and Comparative Fungal Genomics Platform (CFGP, http://cfgp.snu.ac.kr/) (Choi et al., 2013) (Table 1). Protein sequences were aligned by ClustalW, and phylogenetic tree was constructed using the neighbor-joining method in MEGA program (Tamura et al., 2013). Phylogenetic relationships were tested with a bootstrap method using 2,000 repetitions.
Genomic DNA was isolated using two different methods depending on experimental purpose. Genomic DNA for double-joint PCR and Southern blot was isolated according to a standard protocol (Sambrook and Russell, 2001). Genomic DNA used for gene deletion mutant screening by PCR was prepared using a quick extraction method (Chi et al., 2009b). Gene knockout constructs were produced using double-joint PCR (Yu et al., 2004), which fuses three different DNA fragments (two 1 to 1.5 kb fragments corresponding to the 5′- and 3′-flank regions of each target gene and a 2.1 kb fragment containing the Hygromycin B phosphotransferase gene) (Table 2). Resulting constructs were introduced into protoplasts of KJ201 as previously described (Sweigard et al., 1992). Hygromycin B-resistant transformants were screened first using PCR and confirmed by Southern analysis (Sambrook and Russell, 2001) (Fig. 2). Hybridization probes were prepared using the Rediprime™ II DNA Labeling System (GE Healthcare, USA) according to the manufacturer’s instructions, and signals on the membranes were detected using Phosphorimager (BAS-2040, Fuji Photo Film, Japan). For genetic complementation, an intact copy of the mutated gene was amplified by PCR and introduced into protoplasts of individual mutants using a geneticin-resistance gene as a selection marker.
To evaluate mycelial growth, the diameter of colonies grown on MCA and MMA was measured at 9 days post inoculation (dpi) with three replicates. The ability of asexual reproduction was assessed by counting the number of conidia harvested with 5 ml of sterilized distilled water from 6-day-old V8A cultures using a hemacytometer. The size of individual conidia was measured using a light microscope, and conidial germination and appressorium formation were evaluated on a hydrophobic coverslip. Three drops (40 μl per drop) of conidial suspension (approximately 2 to 5 × 104 conidia/ml) on a coverslip were placed in a container with wet paper towel and incubated at 25°C for 4 and 24 h. The proportions of germinating conidia and those forming the appressorium were microscopically counted (at least 100 conidia per replicate with three replicates). To visualize dead conidia, phloxine B was added to conidial suspensions harvested from two-week-old V8A cultures at the final concentration of 20 μg/ml and stained for 1 h. To determine the number of cells in conidia, conidia harvested from 30-day-old OMA cultures were examined using a microscope (at least 500 conidia per replicate with three replicates). To visualize conidial cell wall and nuclei, calcofluor white and Hoechst 33342, respectively were used. To observe the septum formation in hyphae, conidia were inoculated on 50 to 100 μl drop of diluted complete agar (0.6 g of yeast extract, 0.6 g of Casamino acids, 1 g of sucrose, and 15 g of agar per liter) on a slide glass or petri dish. After 36 to 48 hours of incubation at 25°C under humid condition, agar drops were dried in a clean bench and observed using a light microscope. Hyphal morphology was observed using a light microscope after culturing conidia in liquid diluted CM (0.6 g of yeast extract, 0.6 g of Casamino acids, 1 g of sucrose per liter) for 2 days.
To assess the sensitivity to cell cycle inhibitors, conidial suspension (approximately 2 to 5 × 104 conidia/ml) was dropped on hydrophobic coverslip with 0.5 mM hydroxyurea and 2 μg/ml benomyl. Benomyl was dissolved in 95% EtOH to make a 1 mg/ml stock solution. Conidial germination and appressorium formation were microscopically evaluated at 24 hours post inoculation (hpi). For high or low temperature stresses, conidial suspension (approximately 2 to 5 × 104 conidia/ml) was dropped on hydrophobic coverslip, followed by incubation at 37°C and 4°C. After 24 hpi, the samples were moved to 25°C for further growth. Conidial germination and appressorium formation were evaluated at 48 hpi. Physiological response to various chemicals that cause stress was evaluated using a novel colorimetric measurement method using a pH indicator (J. Park et al. manuscript in preparation). Conidial suspension (approximately 1 × 106 conidia/ml) was inoculated in wells of 384-well microtiter plate containing the following chemical solutions in water and incubated at 25°C for 1 day: 100 mM NaCl, 150 mM KCl, 150 mM MgCl2, 100 mM LiCl, 100 mM CaCl2, 0.2 mM EDTA, 5 mM Caffeine, 0.2 mM MnCl2, 0.08 mM ZnSO4, 25 mM hydroxyurea, 5 μg/ml benomyl, 0.2 μM cycloheximide, 1.3 mM 3-AT, 133 ppm calcofluor white, 0.002% SDS, 0.5 mM DTT, 0.0001% methyl viologen, 1 mM H2O2, 40 ppm p-coumaric acid, 13 mM KClO3, and 0.4 mM NaNO2. After the incubation, an alkalinization-inducing medium (L-aspartic acid solution at the final concentration of 2.5 mM with 0.008% bromocresol purple) was added to individual wells to monitor the ambient alkalinization process. Initial pH of the alkalinization-inducing medium was adjusted to be 4.6 by adding NaOH solution. Color change of bromocresol purple (from yellow to purple) was continuously measured for 24 h at every 20 minutes using a microplate reader (Epoch, BioTek, USA) at 589 nm and 432 nm. The relative activity of alkalinization was used as an indirect indicator for physiological response of each strain to individual chemicals.
Conidia were harvested from 7 to 14-day-old V8A cultures, and 10 ml of conidial suspension (approximately 5 × 104 conidia/ml) containing 250 ppm Tween 20 was sprayed onto susceptible rice seedlings (Oryza sativa cv. Nakdongbyeo) at the 4 to 5 leaf stage. The inoculated plants were kept in a dew chamber at 25°C for 24 hours under darkness and then moved to a growth chamber with 16-h-light/8-h-dark cycle. Diseased leaves were evaluated at 6 to 7 dpi. Penetration assays were performed using rice sheaths. Conidial suspension (approximately 2 × 104 conidia/ml) was injected in rice sheath and incubated at 25°C under high humidity. Invasive growth of M. oryzae strains was observed at 48 hpi using a light microscope.
Species in the phyla Ascomycota and Basidiomycota carry 3 to 6 putative FOX TF genes with M. oryzae containing four (Table 1), according to data archived in the Fungal Transcription Factor Database (Park et al., 2008). To study how the M. oryzae FOX TFs have evolved and are related to previously characterized fungal FOX TFs, a phylogenetic analysis was conducted using FOX TFs in M. oryzae and 19 other species (Fig. 1). MGG_06258.7, a M. oryzae FOX TF belonging to Ascomycota clade 1, was closely related to S. cerevisiae FKH1 and FKH2, S. pombe Fkh2, and C. albicans CaFKH2, while MGG_06422.7 is orthologous to S. cerevisiae HCM1 and S. pombe Mei4 and Sep1 (Ascomycota clade 2). Accordingly, these proteins were named as MoFKH1 and MoHCM1, respectively. The remaining two FOX TFs in M. oryzae (MGG_01853.7 and MGG_14496.7) belong to Pezizomycotina-specific clades and were named MoFOX1 and MoFOX2, respectively. MoFOX1 is an ortholog of A. nidulans FhpA, which seems to be a regulator of sexual reproduction (Lee et al., 2005).
These M. oryzae FOX TF genes were mutagenized via transformation-mediated gene replacement to study their roles. Screening of 48, 48, 287, and 1,378 Hygromycin B-resistant transformants with mutant alleles of MoFKH1, MoHCM1, MoFOX1, and MoFOX2, respectively, resulted in 2, 4, 2, and 0 desired mutants. We used one mutant for each gene, including ΔMofkh1-1, ΔMohcm1-4, and ΔMofox1-1, for phenotypic characterizations (Fig. 2 and Table 3). MoFOX2 may be an essential gene given the failure of obtaining a mutant even after screening 1,378 transformants from three independent experiments.
ΔMofkh1 and ΔMohcm1 showed slightly reduced growth on MCA and MMA (Fig. 3A) as well as V8A and OMA (data not shown). The amount of conidia produced by ΔMofkh1 and ΔMohcm1 was similar to that of wild-type strain KJ201 (Fig. 3B), whereas conidial germination was decreased in these mutants (Fig. 3C). Approximately 70% and less than 90% of ΔMofkh1 and ΔMohcm1 conidia, respectively, germinated after 24 hours of incubation. Their reduced conidial germination was not complemented by additional nutrients or longer incubation (data not shown). To test if the reduced germination was due to lower conidial viability, we stained conidia with phloxine B, which accumulates in dead conidia. 10.8% of ΔMofkh1 conidia appeared dead, while ΔMohcm1 and KJ201 produced 0.5 to 1.9% of stillbirth conidia (Fig. 3D), suggesting that the reduced conidial germination in ΔMofkh1 and ΔMohcm1 is not simply due to increased production of nonviable conidia. Growth retardation and germination defect in ΔMofkh1 and ΔMohcm1 were fully restored when an intact copy of the mutated gene was reintroduced. ΔMofox1 was indistinguishable from wild-type in growth and conidial production and germination.
Based on previous studies suggesting the involvement of the orthologs of MoFKH1 and MoHCM1 in cell cycle regulation in model yeasts (Pramila et al., 2006; Zhu et al., 2000), sensitivity of all three mutants to hydroxyurea and benomyl, cell cycle inhibitors, was measured. Hydroxyurea is a reversible inhibitor of ribonucleotide reductase and arrests the initiation and elongation of DNA replication. Benomyl causes spindle depolymerization, thus inhibiting chromosome segregation. While over 90% of germinated-conidia formed the appressorium at 24 hpi in all strains (Fig. 3E), in the presence of hydroxyurea, many more germinated-conidia failed to form appressoria in ΔMohcm1 compared to other strains (Fig. 4A). In the presence of benomyl, the percentage of appressorium formation was reduced to 52.2% in ΔMohcm1, while ≥70% of them still formed the appressorium in other strains (Fig. 4B). Sensitivity of a complemented strain of ΔMohcm1 to these compounds was comparable to that of KJ201.
ΔMofkh1 produced relatively smaller conidia than those produced by KJ201 (Fig. 5A and 5B). However, while KJ201 mainly produced typical three-celled conidia, >10% of ΔMofkh1 conidia contain ≥4 cells (Fig. 6A). The number of abnormal conidia increased as the culture aged with 15.9%, 2.3%, and 0.3% of the conidia containing four, five, and six cells, respectively after 30 days of incubation on OMA (Fig. 6B). Staining with Hoechst 33342 showed that the abnormal conidia of ΔMofkh1 also exhibited an increased number of nuclei (Fig. 6C), suggesting improper control of cell division in this mutant. In addition, ΔMofkh1 mycelia exhibited increased septation (Fig. 7A). When conidia in diluted CM were incubated at 25°C for 2 days, uneven shaped hyphae formed by ΔMofkh1 (Fig. 7B). Imaging by scanning electron microscope revealed highly increased septum formation in this uneven shaped hyphae (Fig. 7C). The complemented strain Mofkh1c and other mutant strains (ΔMohcm1 and ΔMofox1) displayed wild-type hyphal morphology and septation.
To study if and how individual mutations affect physiological response to various agents that cause stress, external pH change, as an indirect indicator of the physiological status of cells, was monitored via the use of a colorimetric measurement method (J. Park et al., manuscript in preparation) and compared with change in KJ201 (Table 4). ΔMofkh1 showed alkalinization activity significantly different from KJ201 in the presence of salts (NaCl, KCl, and LiCl), heavy metal (ZnSO4), cell cycle inhibitor (benomyl), a plant phenolic compound (p-coumaric acid), and DTT that causes endoplasmic reticulum (ER) stress. The alkalinization activity of ΔMohcm1 and ΔMofox1 was comparable to that of KJ201 under all conditions.
To evaluate their sensitivity to high and low temperatures, we treated conidia of KJ201 and its mutants at 37°C and 4°C for 24 hours. Germination and appressorium formation rates of ΔMofkh1 conidia were reduced from 72.7% to 43% and 94.5% to 34.5%, respectively after heat treatment, while other strains showed slight or no reduction (Fig. 8A). Although the number of dead conidia in ΔMofkh1 slightly increased after the heat treatment, >80% of them were still viable. Other strains including KJ201 showed less than 3% of stillbirth conidia after the heat treatment (Fig. 8B). All the mutant strains were not sensitive to the cold shock.
To evaluate virulence, two methods were employed. Upon spray inoculation of susceptible rice seedlings with conidia, all three mutants and KJ201 caused lesions on rice leaves. However, ΔMofkh1 formed relatively smaller lesions than the others. Most lesions formed by ΔMofkh1 remained dark-brown spot and expanded more slowly compared to those caused by KJ201 (Fig. 9A). Invasive growth of ΔMofkh1 in rice sheath cells also was slightly reduced compared to that of KJ201 (Fig. 9B). Genetic complementation confirmed that loss of MoFKH1 was responsible for reduced virulence.
TFs are pivotal components in controlling and coordinating diverse biological processes by orchestrating gene expression in response to various stimuli and developmental cues. Although well-annotated genome sequences have accelerated functional analyses of TFs in M. oryzae (Choi et al., 2009; Guo et al., 2011; Kim et al., 2009; Mehrabi et al., 2008; Park et al., 2008; Park et al., 2013), most predicted TFs still remain to be characterized. The focus of this study was on determining the roles of the FOX TF gene family, which consists of four genes, in controlling development, pathogenicity, and stress response.
Species in Ascomycota and Basidiomycota contain 3-6 FOX TF genes, whereas species in the phylum Zygomycota carry 10 or more (e.g., 10, 11, and 12 genes in Mucor circinelloides, Phycomyces blakesleeanus, and Rhizopus oryzae, respectively) (Park et al., 2008). Phylogenetic analysis using characterized and predicted fungal FOX TFs revealed multiple seemingly phylum- or subphylum-specific clades. The number variation and taxon-specific distribution of members of this gene family suggest its dynamic evolution probably driven by unique regulatory needs in individual taxa.
MoFKH1 is a homolog of S. cerevisiae FKH1 and FKH2 (Zhu et al., 2000), S. pombe Fkh2 (Bulmer et al., 2004), and C. albicans CaFKH2 (Bensen et al., 2002). These proteins participate in controlling cell cycle, cell separation, and/or morphogenesis. Since septation is linked to cell cycle progression, abnormal septation observed in ΔMofkh1 in conidia and mycelia (Figs. 6 and 7) suggest the involvement of MoFKH1 in cell cycle control. Increased number of nuclei in ΔMofkh1 conidia and its susceptibility to benomyl also support this supposition. Although conidial germination and appressorium formation was not hindered by benomyl, the alkalinization activity in ΔMofkh1 was significantly reduced compared to that of KJ201, suggesting that loss of MoFKH1 affects the regulation of certain physiological processes. Results from this and previous studies (Bulmer et al., 2004; Zhu et al., 2000) suggest that the function of MoFKH1 and its orthologs is conserved within Ascomycota. Reduced germination rate of ΔMofkh1 conidia may be due to a combination of increased production of stillbirth conidia and unknown defects in controlling germ tube emergence. Park et al. (2013) reported that expression of MoFKH1 increased in conidia, supporting its role in controlling the production of conidia.
As demonstrated in this study (Table 4), the use of ambient pH change as an indicator for monitoring physiological change in response to various agents that cause cellular stress offers a novel means for studying gene function. Given that many physiological processes affect ambient pH, this assay alone is not sufficient for revealing the nature of physiological change(s) caused by individual mutations, but provides useful clues to recognizing potential defects. In addition, because it employs microtiter plates and a plate reader, mutant responses to multiple stimuli can be quickly assayed. Reduced ambient alkalinization in the presence of ionic stresses (NaCl, KCl, LiCl, and ZnSO4) suggests a role of MoFKH1 in controlling cellular processes affected by these stresses. Higher sensitivity of ΔMofkh1 to DTT and heat shock also indicated that the function of MoFKH1 might be related to the unfolded protein response. Compared to Δdes1 (Chi et al., 2009a), which drastically affected alkalinization activity in response to H2O2 (data not shown), ΔMofkh1 was indistinguishable from KJ201 under that condition. This suggests that reduced virulence of ΔMofkh1 may be caused by defects other than its reduced ability to manage oxidative stress. One possible defect is unstable cell cycle progression. Several studies indicated the connection of orderly cell cycle control to proper development and pathogenicity in M. oryzae. SEP1, a putative homolog of Cdc7 in S. pombe, is essential for positioning of cell division sites, and its temperature-sensitive mutant showed reduced virulence (Saunders et al., 2010). MoCDC15, a homolog of Cdc15 in S. pombe, is important for growth, septation, asexual development and pathogenicity (Goh et al., 2011). Expression of MoCDC15 was elevated in ΔMofkh1 (Goh et al., 2011), suggesting that MoFKH1 controls cell cycle control by modulating the expression of MoCDC15. When the phenotypes of ΔMofkh1 and ΔMoCDC15TDNA are compared, they displayed similar reduction in mycelial growth, abnormal septation, reduced conidial germination, increased number of dead conidia, and reduced pathogenicity. The reduced alkalinization activity of ΔMofkh1 in response to a plant phenolic compound (p-coumaric acid) suggests that reduced virulence in ΔMofkh1 is in part caused by defect in responding to plant chemical defense.
MoHCM1 is homologous to S. cerevisiae HCM1 (Pramila et al., 2006), S. pombe Sep1 (Ribár et al., 1999) and Mei4 (Horie et al., 1998). Benomyl sensitivity of ΔMohcm1 was similar to that of the S. cerevisiae HCM1 mutant, suggesting their involvement in regulating conserved cell cycle-related processes. Given that ΔMohcm1 normally formed septa, its role in cell cycle control may be distinct from that of MoFKH1. MoHCM1 was required for normal germination and growth but dispensable for virulence and several stress responses. The failure in disrupting ΔMoFOX2 may be due to low frequency of homologous recombination at this locus or lethality of resulting mutants. MoFOX1 is an ortholog of A. nidulans FhpA, a predicted regulator of sexual reproduction. However, we could not test its role in sexual reproduction because M. oryzae KJ201 is infertile. Loss of MoFOX1 did not cause noticeable defects in development, pathogenicity, and all of tested stress conditions. Taken together, results from this study contribute to uncovering the transcriptional regulatory network controlling development, pathogenicity, and stress response in M. oryzae.
Acknowledgments
This work was supported by grants from the National Research Foundation of Korea (2008-0061897 and 2013-003196), and the Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ00821201) from the Rural Development Administration, Republic of Korea. Park, J., Kong, S., and Kim, S. are grateful for a graduate fellowship from the Brain Korea 21 Program.
Fig. 1.
Phylogenetic analysis of putative FOX TFs encoded by selected species in Ascomycota and Basidiomycota. A phylogenetic tree based on their protein sequences was constructed using the neighbor-joining method with bootstrap analysis (2,000 replicates). Each number next to branches denotes the bootstrap value. Functionally characterized proteins and four putative FOX TFs in M. oryzae (MoFKH1, MoHCM1, MoFOX1, and MoFOX2) were noted with black arrows. Phylum and subphylum-specific clades in Ascomycota were marked by black lines.
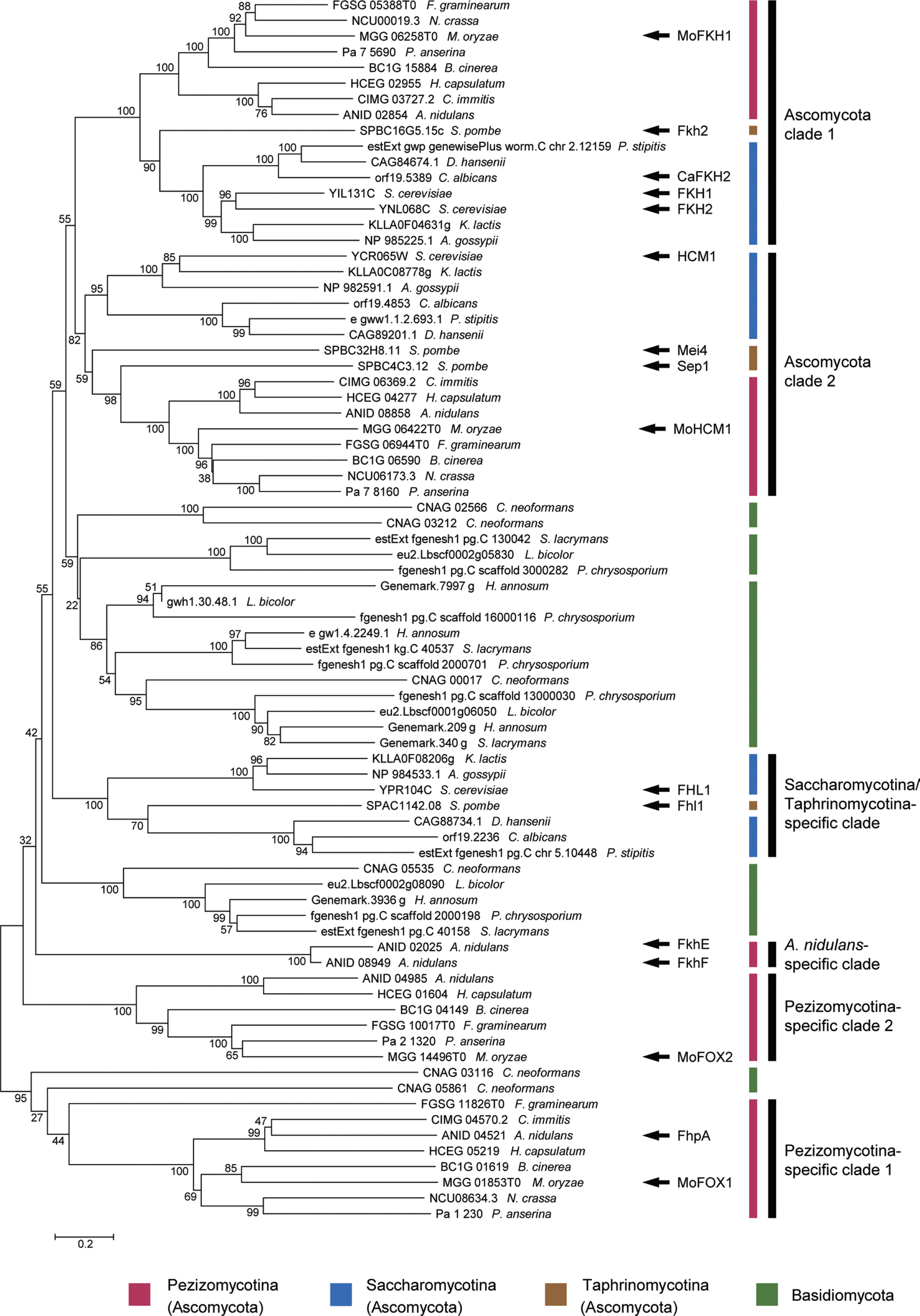
Fig. 2.
Gene deletion strategy. M. oryzae MoFKH1, MoHCM1, and MoFOX1 were deleted via transformation-mediated targeted gene replacement. Southern results confirmed gene deletion.
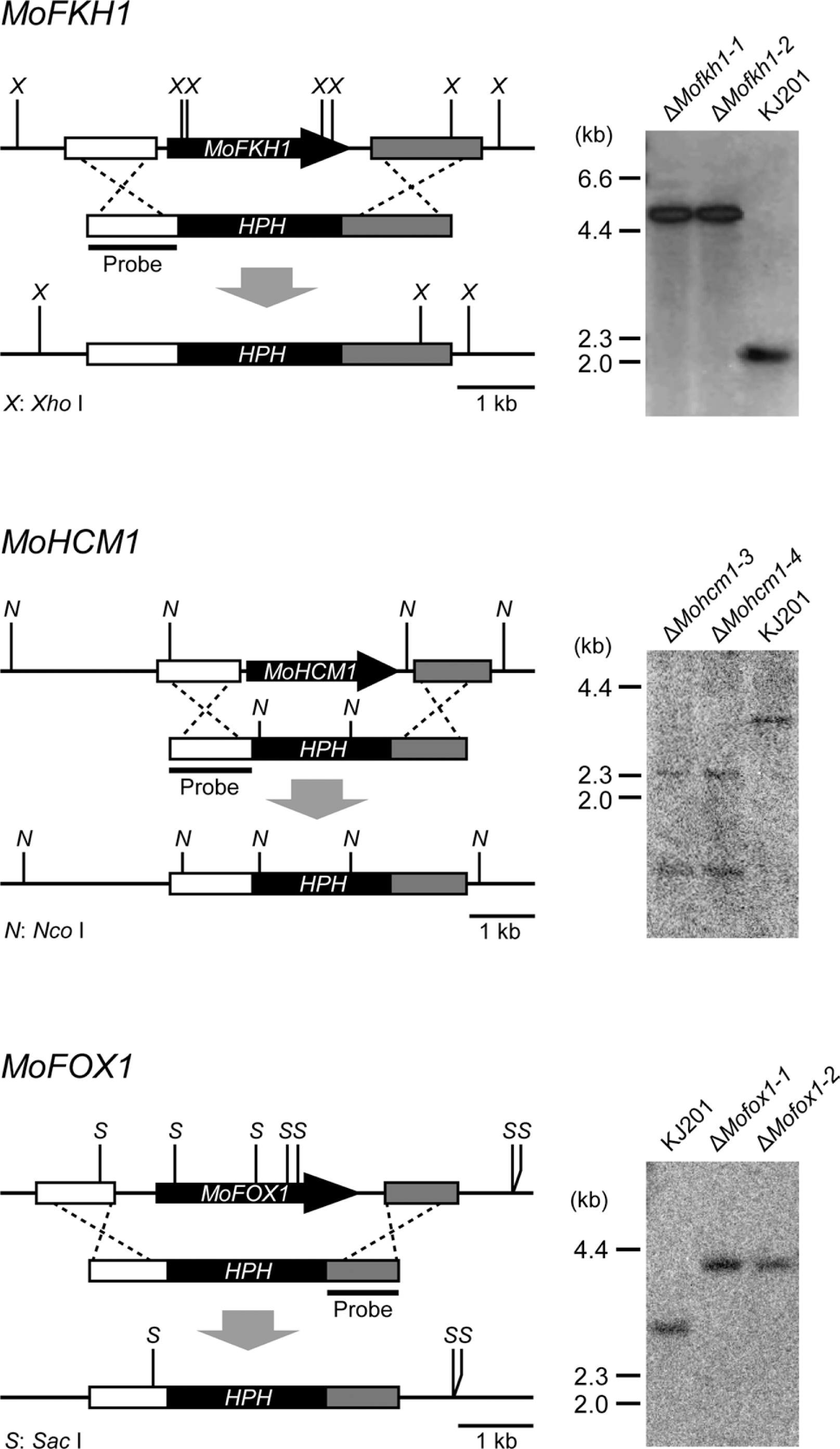
Fig. 3.
Colonial growth, spore production and germination, and appressorium formation. (A) Colony diameter of wild-type (KJ201), mutants (ΔMofkh1, ΔMohcm1, and ΔMofox1) and complemented mutant strains (Mofkh1c and Mohcm1c) on MCA and MMA was measured at 9 dpi. (B) The numbers of conidia produced by the same strains are graphically presented. (C) Conidial germination at 4 hpi and 24 hpi is shown. (D) Percentages of dead conidia are shown. (E) Percentages of germinated conidia that formed the appressorium at 24 hpi are shown.
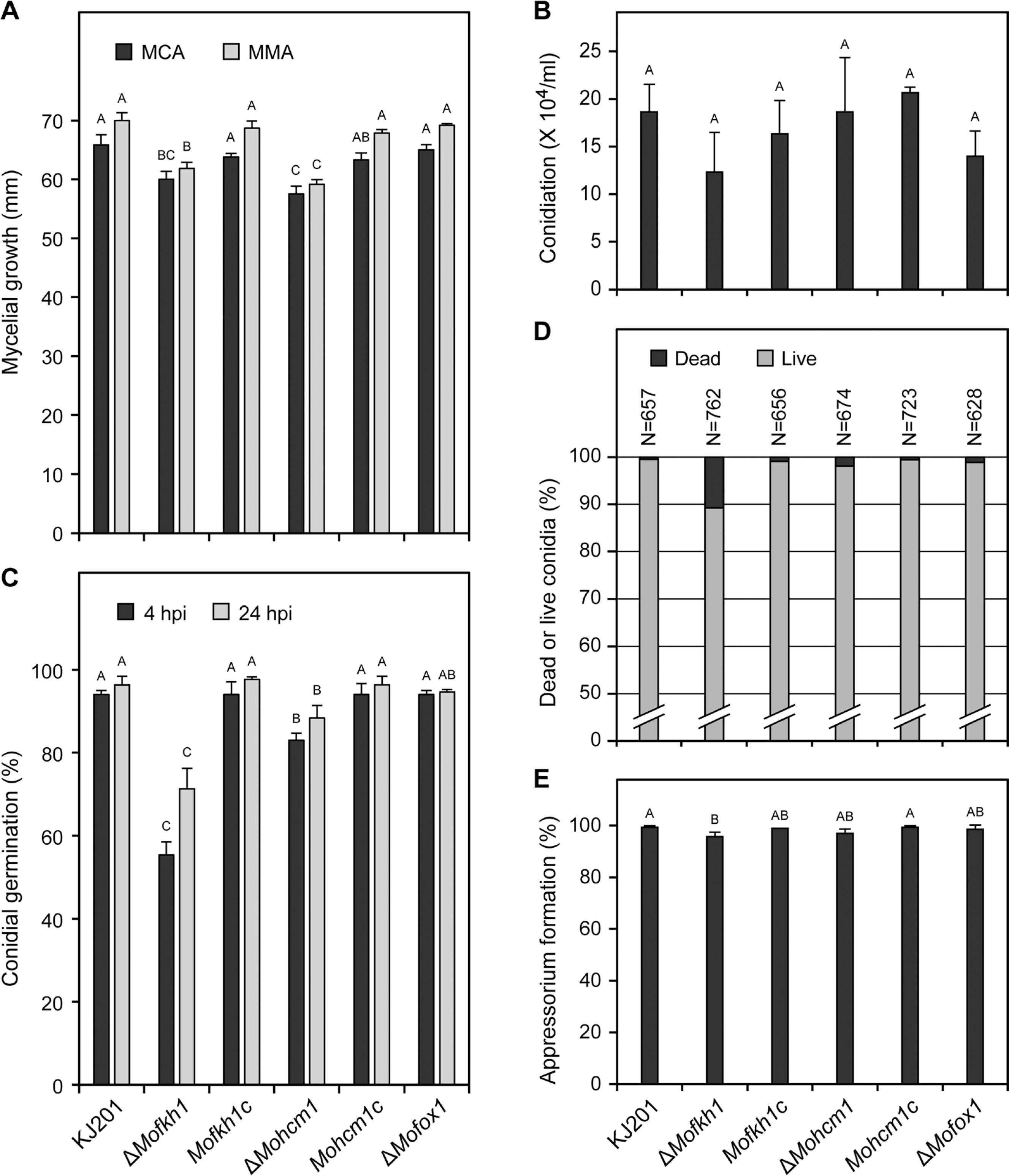
Fig. 4.
Effect of cell cycle inhibitors on conidial germination and appressorium formation. Conidial germination and appressorium formation in the presence of (A) hydroxyurea (0.5 mM) and (B) benomyl (2 μg/ml with 0.19% EtOH) are shown. The control for benomyl treatment was 0.19% EtOH.

Fig. 5.
Effect of MoFKH1 deletion on conidial size. (A) The average length and width of conidia produced by KJ201, mutants (ΔMofkh1, ΔMohcm1, and ΔMofox1) and complemented mutant strains (Mofkh1c and Mohcm1c) are shown. (B) Distribution patterns of conidial length and width among 100 conidia each from KJ201, ΔMofkh1 and Mofkh1c are shown.
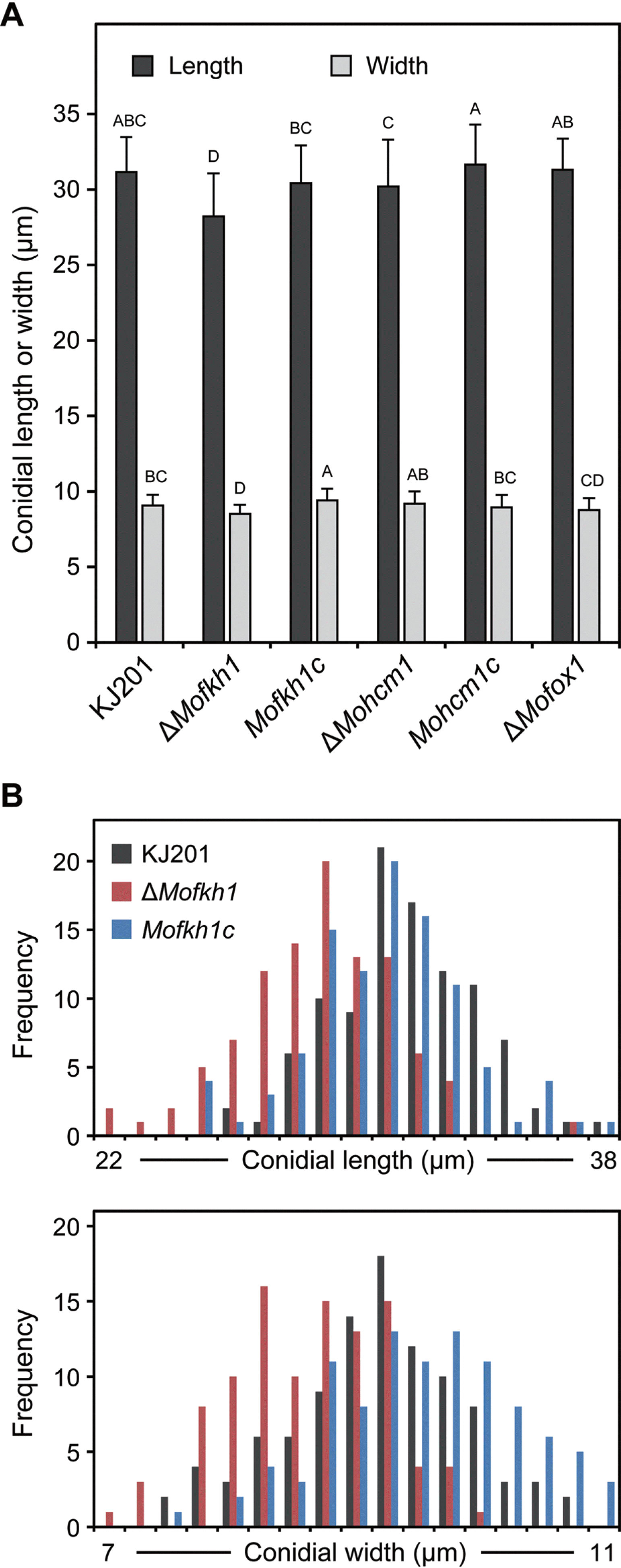
Fig. 6.
Changes in septation and cell number by ΔMofkh1. (A) Septation in KJ201 and ΔMofkh1 conidia was visualized by calcofluor white staining. Bars = 10 μm. (B) Increased cell numbers in ΔMofkh1 conidia. (C) Nuclei in KJ201 and ΔMofkh1 conidia were stained using Hoechst 33342. White arrows indicate nuclei. Bars = 10 μm.
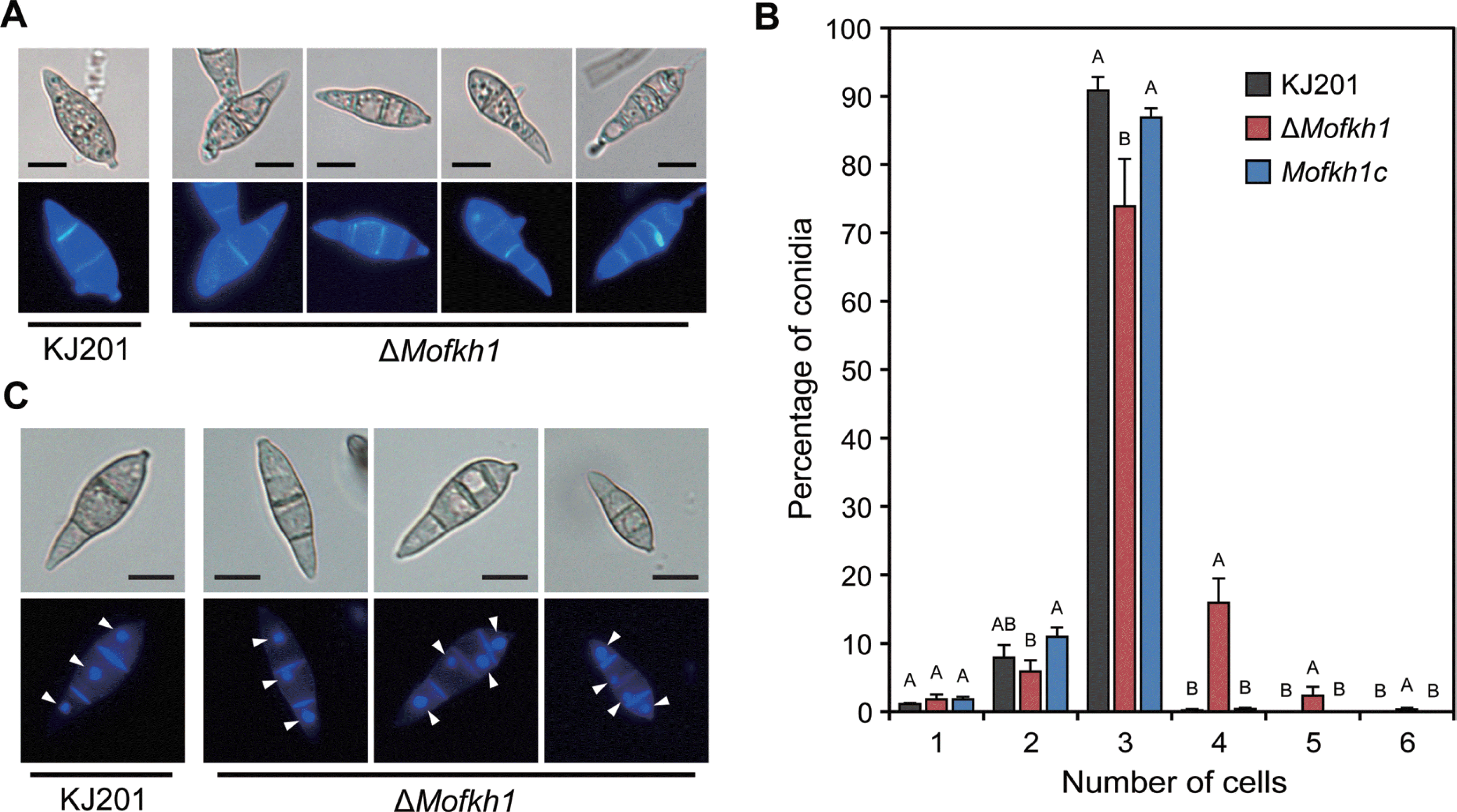
Fig. 7.
Abnormal septation and morphology in ΔMofkh1 mycelia. (A) ΔMofkh1 mycelia showed increased septation, which was denoted by black arrows. (B) Abnormal mycelial shapes of ΔMofkh1 observed using a light microscope and (C) scanning electron micrographs are shown. Septa are marked using white arrows.
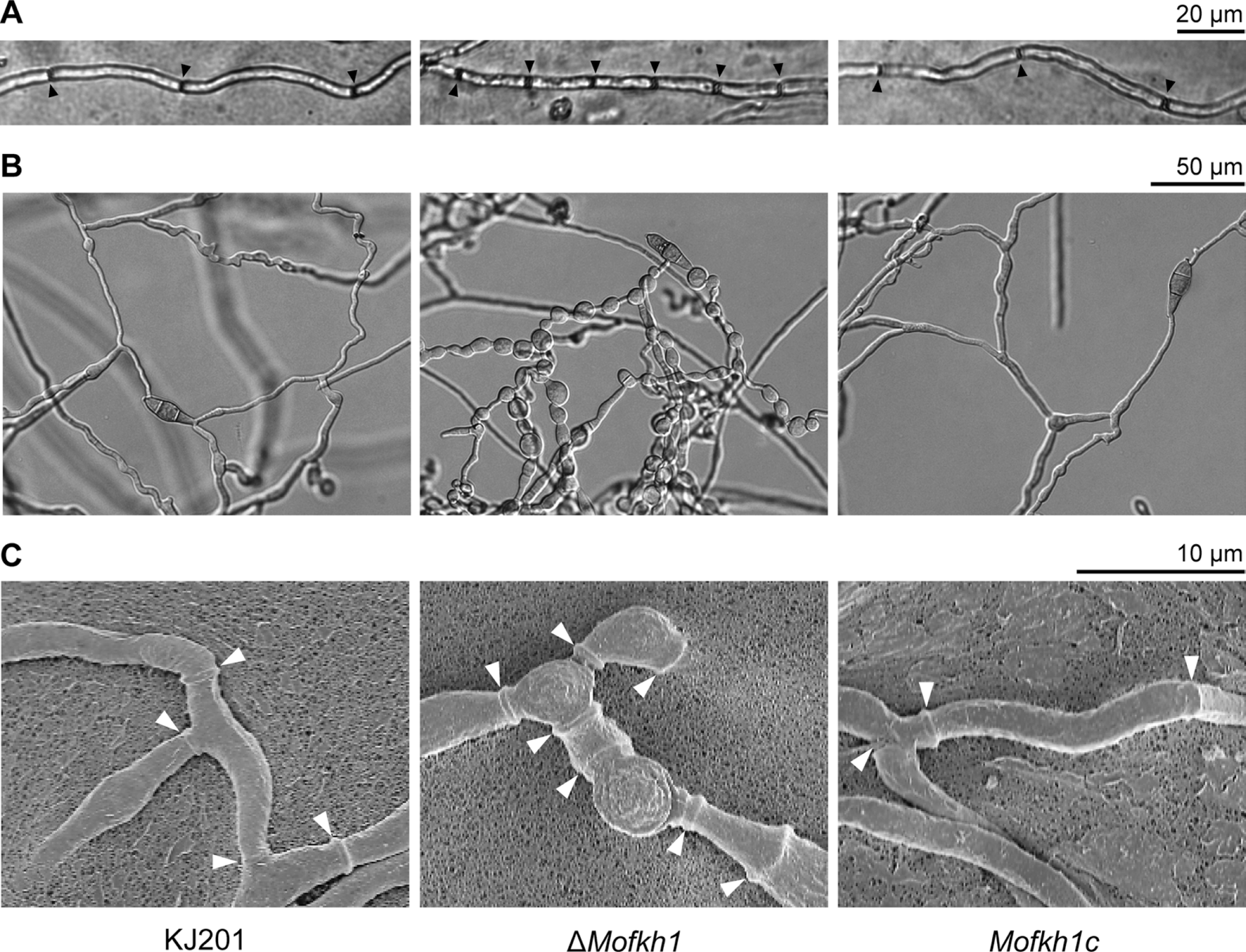
Fig. 8.
Responses of conidia of FOX TF mutants to temperature stresses. (A) Conidial germination and appressorium formation after high (37°C) and low (4°C) temperature treatments are shown. (B) Percentages of dead conidia after harvest (left panel) and following an incubation at 37°C for 24 h (right panel) are shown.
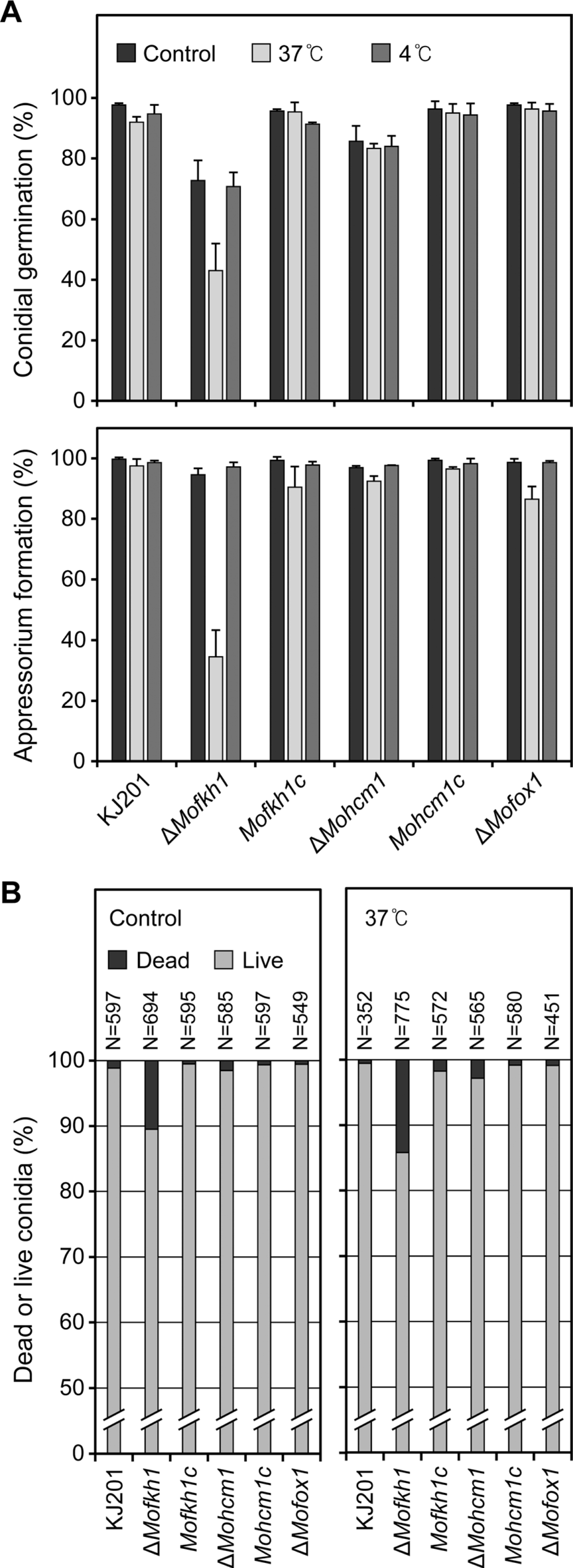
Fig. 9.
Reduced symptoms on rice leaves and infectious growth in rice sheath cells by ΔMofkh1. (A) Lesions formed by KJ201, ΔMofkh1, and Mofkh1c on rice leaves after spray inoculation are shown. (B) Invasive growth of KJ201, ΔMofkh1, and Mofkh1c in rice sheath at 48 hpi. Bars = 50 μm.

Table 1.
Number of FOX TFs in the fungal species shown in Fig. 1
Table 2.
Primers used for producing gene knockout constructs
Table 3.
M. oryzae strains used in this study
Table 4.
Relative alkalinization activity of FOX TF mutants under various conditions
| Treatments |
Relative alkalinization activity of each straina
|
|||||
|---|---|---|---|---|---|---|
| KJ201 | ΔMofkh1 | Mofkh1c | ΔMohcm1 | Mohcm1c | ΔMofox1 | |
| NaCl | 1.00A | 0.73 ± 0.04B | 1.06 ± 0.09A | 1.03 ± 0.03A | 1.12 ± 0.08A | 0.98 ± 0.11A |
| KCl | 1.00A | 0.81 ± 0.11B | 1.02 ± 0.05A | 1.03 ± 0.04A | 1.10 ± 0.03A | 0.94 ± 0.11AB |
| MgCl2 | 1.00A | 1.04 ± 0.19A | 1.00 ± 0.02A | 1.04 ± 0.05A | 1.10 ± 0.13A | 0.90 ± 0.12A |
| LiCl | 1.00A | 0.69 ± 0.09B | 1.01 ± 0.07A | 1.10 ± 0.09A | 1.12 ± 0.08A | 0.98 ± 0.14A |
| CaCl2 | 1.00A | 1.04 ± 0.21A | 1.00 ± 0.05A | 1.04 ± 0.07A | 1.12 ± 0.12A | 0.93 ± 0.08A |
| EDTA | 1.00AB | 0.85 ± 0.15B | 0.98 ± 0.19AB | 1.09 ± 0.06AB | 1.10 ± 0.09AB | 1.22 ± 0.08A |
| Caffeine | 1.00AB | 0.95 ± 0.03B | 0.99 ± 0.12B | 1.21 ± 0.05A | 1.10 ± 0.08AB | 1.05 ± 0.10AB |
| MnCl2 | 1.00A | 0.78 ± 0.08A | 1.05 ± 0.17A | 1.03 ± 0.11A | 1.16 ± 0.11A | 0.87 ± 0.30A |
| ZnSO4 | 1.00A | 0.58 ± 0.14B | 0.98 ± 0.09A | 0.96 ± 0.06A | 1.13 ± 0.15A | 0.92 ± 0.20A |
| Hydroxyurea | 1.00AB | 0.78 ± 0.10B | 1.07 ± 0.06A | 1.14 ± 0.06A | 1.14 ± 0.09A | 1.00 ± 0.20AB |
| Benomyl | 1.00A | 0.59 ± 0.13B | 1.01 ± 0.09A | 1.02 ± 0.05A | 1.14 ± 0.08A | 0.92 ± 0.10A |
| Cycloheximide | 1.00AB | 1.18 ± 0.11A | 0.94 ± 0.03B | 1.00 ± 0.06AB | 1.06 ± 0.06AB | 0.85 ± 0.14B |
| 3-Amino-1,2,4-triazole | 1.00A | 0.96 ± 0.08A | 1.03 ± 0.16A | 1.07 ± 0.05A | 1.02 ± 0.01A | 0.97 ± 0.14A |
| Calcofluor white | 1.00A | 1.14 ± 0.09A | 0.98 ± 0.13A | 1.07 ± 0.07A | 0.96 ± 0.06A | 1.04 ± 0.12A |
| SDS | 1.00A | 0.76 ± 0.11A | 1.06 ± 0.26A | 1.24 ± 0.25A | 1.22 ± 0.16A | 1.22 ± 0.30A |
| DTT | 1.00A | 0.32 ± 0.05B | 1.25 ± 0.30A | 1.27 ± 0.10A | 1.19 ± 0.16A | 1.06 ± 0.25A |
| Methyl viologen | 1.00AB | 0.81 ± 0.13B | 1.10 ± 0.08A | 1.18 ± 0.08A | 1.09 ± 0.12A | 0.81 ± 0.09B |
| H2O2 | 1.00A | 1.03 ± 0.22A | 1.02 ± 0.12A | 1.19 ± 0.20A | 1.04 ± 0.14A | 0.89 ± 0.17A |
| p-Coumaric acid | 1.00A | 0.74 ± 0.05B | 1.06 ± 0.08A | 1.00 ± 0.05A | 1.06 ± 0.10A | 0.92 ± 0.05A |
| KClO3 | 1.00A | 0.98 ± 0.11A | 1.07 ± 0.07A | 1.10 ± 0.04A | 1.05 ± 0.04A | 0.92 ± 0.16A |
| NaNO2 | 1.00A | 0.98 ± 0.12A | 1.07 ± 0.03A | 1.10 ± 0.10A | 1.03 ± 0.03A | 0.96 ± 0.10A |
a Relative alkalinization activity = (alkalinization activity of mutant in stress condition/alkalinization activity of mutant in NT)/(alkalinization activity of KJ201 in stress condition/alkalinization activity of KJ201 in NT)
References
Bensen, ES, Filler, SG and Berman, J 2002. A forkhead transcription factor is important for true hyphal as well as yeast morphogenesis in Candida albicans. Eukaryot. Cell. 1:787-798.


Bulmer, R, Pic-Taylor, A, Whitehall, SK, Martin, KA, Millar, JBA, Quinn, J and Morgan, BA 2004. The forkhead transcription factor Fkh2 regulates the cell division cycle of Schizosaccharomyces pombe. Eukaryot. Cell. 3:944-954.


Carlsson, P and Mahlapuu, M 2002. Forkhead transcription factors: key players in development and metabolism. Dev Biol. 250:1-23.
Chi, M-H, Park, S-Y, Kim, S and Lee, Y-H 2009a. A novel pathogenicity gene is required in the rice blast fungus to suppress the basal defenses of the host. PLoS Pathog. 5:e1000401.
Chi, M-H, Park, S-Y and Lee, Y-H 2009b. A quick and safe method for fungal DNA extraction. Plant Pathol J. 25:108-111.
Choi, J, Cheong, K, Jung, K, Jeon, J, Lee, G-W, Kang, S, Kim, S, Lee, Y-W and Lee, Y-H 2013. CFGP 2.0: a versatile web-based platform for supporting comparative and evolutionary genomics of fungi and Oomycetes. Nucleic Acids Res. 41:D714-D719.
Choi, J, Kim, Y, Kim, S, Park, J and Lee, Y-H 2009. MoCRZ1, a gene encoding a calcineurin-responsive transcription factor, regulates fungal growth and pathogenicity of Magnaporthe oryzae. Fungal Genet Biol. 46:243-254.

Dean, RA, Talbot, NJ, Ebbole, DJ, Farman, ML, Mitchell, TK, Orbach, MJ, Thon, M, Kulkarni, R, Xu, J-R, Pan, H, Read, ND, Lee, Y-H, Carbone, I, Brown, D, Oh, YY, Donofrio, N, Jeong, JS, Soanes, DM, Djonovic, S, Kolomiets, E, Rehmeyer, C, Li, W, Harding, M, Kim, S, Lebrun, M-H, Bohnert, H, Coughlan, S, Butler, J, Calvo, S, Ma, L-J, Nicol, R, Purcell, S, Nusbaum, C, Galagan, JE and Birren, BW 2005. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature. 434:980-986.

Ebbole, DJ 2007. Magnaporthe as a model for understanding host-pathogen interactions. Annu Rev Phytopathol. 45:437-456.

Goh, J, Kim, KS, Park, J, Jeon, J, Park, S-Y and Lee, Y-H 2011. The cell cycle gene MoCDC15 regulates hyphal growth, asexual development and plant infection in the rice blast pathogen Magnaporthe oryzae. Fungal Genet Biol. 48:784-792.

Guo, M, Chen, Y, Du, Y, Dong, Y, Guo, W, Zhai, S, Zhang, H, Dong, S, Zhang, Z, Wang, Y, Wang, P and Zheng, X 2011. The bZIP transcription factor MoAP1 mediates the oxidative stress response and is critical for pathogenicity of the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 7:e1001302.


Hermann-Le Denmat, S, Werner, M, Sentenac, A and Thuriaux, P 1994. Suppression of yeast RNA polymerase III mutations by FHL1, a gene coding for a fork head protein involved in rRNA processing. Mol Cell Biol. 14:2905-2913.


Horie, S, Watanabe, Y, Tanaka, K, Nishiwaki, S, Fujioka, H, Abe, H, Yamamoto, M and Shimoda, C 1998. The Schizosaccharomyces pombe mei4+ gene encodes a meiosis-specific transcription factor containing a forkhead DNA-binding domain. Mol Cell Biol. 18:2118-2129.


Jürgens, G, Wieschaus, E, Nüsslein-Volhard, C and Kluding, H 1984. Mutations affecting the pattern of the larval cuticle in Drosophila melanogaster. II. Zygotic loci on the third chromosome. Roux's Arch Dev Biol. 193:283-295.
Kim, S, Park, J, Park, S-Y, Mitchell, TK and Lee, Y-H 2010. Identification and analysis of in planta expressed genes of Magnaporthe oryzae. BMC Genomics. 11:104.


Kim, S, Park, S-Y, Kim, KS, Rho, H-S, Chi, M-H, Choi, J, Park, J, Kong, S, Park, J, Goh, J and Lee, Y-H 2009. Homeobox transcription factors are required for conidiation and appressorium development in the rice blast fungus Magnaporthe oryzae. PLoS Genet. 5:e1000757.


Koranda, M, Schleiffer, A, Endler, L and Ammerer, G 2000. Forkhead-like transcription factors recruit Ndd1 to the chromatin of G2/M-specific promoters. Nature. 406:94-98.

Kumar, R, Reynolds, DM, Shevchenko, A, Shevchenko, A, Goldstone, SD and Dalton, S 2000. Forkhead transcription factors, Fkh1p and Fkh2p, collaborate with Mcm1p to control transcription required for M-phase. Curr Biol. 10:896-906.

Lee, B-Y, Han, S-Y, Choi, HG, Kim, JH, Han, K-H and Han, D-M 2005. Screening of growth- or development-related genes by using genomic library with inducible promoter in Aspergillus nidulans. J Microbiol. 43:523-528.

Mehrabi, R, Ding, S and Xu, J-R 2008. MADS-box transcription factor Mig1 is required for infectious growth in Magnaporthe grisea. Eukaryot. Cell. 7:791-799.


Park, J, Park, J, Jang, S, Kim, S, Kong, S, Choi, J, Ahn, K, Kim, J, Lee, S, Kim, S, Park, B, Jung, K, Kim, S, Kang, S and Lee, Y-H 2008. FTFD: an informatics pipeline supporting phylogenomic analysis of fungal transcription factors. Bioinformatics. 24:1024-1025.

Park, M-H, Kim, H-Y, Kim, JH and Han, K-H 2010. Gene structure and function of fkhE a forkhead gene in a filamentous fungus Aspergillus nidulans. Kor J Mycol. 38:160-166.
Park, S-Y, Choi, J, Lim, S-E, Lee, G-W, Park, J, Kim, Y, Kong, S, Kim, S, Rho, H-S, Jeon, J, Chi, M-H, Kim, S, Khang, CH, Kang, S and Lee, Y-H 2013. Global expression profiling of transcription factor genes provides new insights into pathogenicity and stress responses in the rice blast fungus. PLoS Pathog. 9:e1003350.


Pramila, T, Wu, W, Miles, S, Noble, WS and Breeden, LL 2006. The Forkhead transcription factor Hcm1 regulates chromosome segregation genes and fills the S-phase gap in the transcriptional circuitry of the cell cycle. Genes Dev. 20:2266-2278.
Ribár, B, Grallert, A, Oláh, É and Szállási, Z 1999. Deletion of the sep1+ forkhead transcription factor homologue is not lethal but causes hyphal growth in Schizosaccharomyces pombe. Biochem Biophys Res Commun. 263:465-474.
Sambrook, J and Russell, DW 2001. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Saunders, DGO, Dagdas, YF and Talbot, NJ 2010. Spatial uncoupling of mitosis and cytokinesis during appressorium-mediated plant infection by the rice blast fungus Magnaporthe oryzae. Plant Cell. 22:2417-2428.


Shimeld, SM, Degnan, B and Luke, GN 2010. Evolutionary genomics of the Fox genes: origin of gene families and the ancestry of gene clusters. Genomics. 95:256-260.
Soanes, DM, Chakrabarti, A, Paszkiewicz, KH, Dawe, AL and Talbot, NJ 2012. Genome-wide transcriptional profiling of appressorium development by the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 8:e1002514.


Sweigard, JA, Chumley, FG and Valent, B 1992. Disruption of a Magnaporthe grisea cutinase gene. Mol Gen Genet. 232:183-190.

Szilagyi, Z, Batta, G, Enczi, K and Sipiczki, M 2005. Characterisation of two novel fork-head gene homologues of Schizosaccharomyces pombe: their involvement in cell cycle and sexual differentiation. Gene. 348:101-109.

Talbot, NJ 1995. Having a blast: exploring the pathogenicity of Magnaporthe grisea. Trends Microbiol. 3:9-16.

Talbot, NJ 2003. On the trail of a cereal killer: exploring the biology of Magnaporthe grisea. Annu Rev Microbiol. 57:177-202.

Talbot, NJ, Ebbole, DJ and Hamer, JE 1993. Identification and characterization of MPG1, a gene involved in pathogenicity from the rice blast fungus Magnaporthe grisea. Plant Cell. 5:1575-1590.


Tamura, K, Stecher, G, Peterson, D, Filipski, A and Kumar, S 2013. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 30:2725-2729.
Weigel, D, Jürgens, G, Küttner, F, Seifert, E and Jäckle, H 1989. The homeotic gene fork head encodes a nuclear protein and is expressed in the terminal regions of the Drosophila embryo. Cell. 57:645-658.

Wilson, RA and Talbot, NJ 2009. Under pressure: investigating the biology of plant infection by Magnaporthe oryzae. Nat Rev Microbiol. 7:185-195.

- TOOLS
-
METRICS

- Related articles
-
Comparative Pathogenicity and Host Ranges of
Magnaporthe oryzae and Related Species2020 August;36(4)



 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print



