Expression of colSR Genes Increased in the rpf Mutants of Xanthomonas oryzae pv. oryzae KACC10859
Article information
Abstract
The rpf genes and colSXOO1207/colRXOO1208 were known to require for virulence of Xanthomonas oryzae pv. oryzae (Xoo). In Xoo KACC10331 genome, two more colS/colR genes, colSXOO3534 (raxH)/colRXOO3535 (raxR) and colSXOO3762/colRXOO3763 were annotated. The colSXOO3534/colRXOO3535 were known to control AvrXa21 activity and functions of colSXOO3762/colRXOO3763 were unknown in Xoo. To characterize the relationship between rpf and colS/colR genes, expression of colS/colR genes in Rpf mutants of Xoo were analyzed with quantitative reverse transcription PCR (qRT-PCR). Expressions of all three colS/colR genes increased in the rpfF mutant in which DSF synthesis is defective. Expression of colSXOO1207/colRXOO1208, colSXOO3534/colRXOO3535 and colSXOO3762/colRXOO3763 increased 2, 2–7, 3–13 folds respectively. Expression of colSXOO3534 and colSXOO3762 also increased 2–4 folds in the rpfG mutant in which the signal from DSF is no longer transferred to down-stream. Expression of the other colS/colR genes was not significantly changed in the rpfG mutant compared to the wild type. Since RpfF and RpfG are responsible for DSF synthesis and signal transfer from DSF to down-stream to regulate virulence gene expression, these results suggest that the DSF and DSF-mediated signal regulate negatively three colS/colR genes in Xoo.
Plant pathogenic bacteria must be virulent and able to suppress host resistance to cause a disease on host plant. As bacterial two-component system (TCS), which perceives outside signals and regulates responses inside bacterial cell, rpfC/rpfG and colSXOO1207/colRXOO1208 are required for virulence of Xanthomonas oryzae pv. oryzae (Xoo) (Slater et al., 2000; Subramoni et al., 2012). The Rpf (regulation of pathogenicity factors) system is known to regulate virulence by the cell-cell communication in X. campestris pv. campestris (Xcc) (Barber et al., 1997; He et al., 2007; Slater et al., 2000; Tang et al., 1991). Among rpf genes that were identified as a cluster (rpfA-I) (Tang et al., 1991), RpfF are responsible for diffusible signal factor (DSF) production (Barber et al., 1997; Wang et al., 2004), and RpfC and RpfG comprise a two-component system, which senses the DSF signal and transfers it to signal cascades involved in virulence (Slater et al., 2000). Virulence factor production and biofilm dispersal are controlled by cyclic di-GMP and Clp, which are on downstream of RpfG (He et al., 2007; Ryan et al., 2007). The rpf genes and functions of the core rpf genes, rpfB, rpfC, rpfF, rpfG, are well conserved in Xoo (Chatterjee and Sonti, 2002; Jeong et al., 2008).
The colS/colR genes, which encode a two-component system, were originally identified from the root-colonizing bacterium Pseudomonas fluorescens and found to be involved in the capacity of the bacterium to colonize plant roots (Dekkers et al., 1998). Subsequently, colS/colR genes were reported to regulate different biological responses to transposition of transposon and various stresses including phenol and heavy metals (Hu and Zhao, 2007; Kivistik et al., 2006). The colS/colR genes are required for virulence of several important plant pathogens including Xoo (Subramoni et al., 2012; Yan and Wang, 2011; Zhang et al., 2008).
Mutation of colSXOO1207/colRXOO1208 decreased virulence of Xoo. These mutations also caused growth defect in iron-limiting condition and deficiency in elielement-citation of hypersensitive response on non-host tomato (Subramoni et al., 2012). Another colS/colR genes, colSXOO3534 (raxH)/colRXOO3535 (raxR), were known to control avrXa21 activity in Xoo pXO99A (Burdman et al., 2004; Lee et al., 2008). These previously published results indicate two virulence regulation systems, Rpf and ColS/ColR, control virulence in Xoo. To characterize the relationship between rpf and colS/colR genes for virulence regulation, expression of colS/colR genes in Rpf mutants of Xoo were analyzed with quantitative reverse transcription PCR (qRT-PCR).
Three colS/colR genes are conserved in the important plant pathogens
In Xoo KACC10331 genome, three colS/colR genes, colSXOO1207/colRXOO1208, colSXOO3534/colRXOO3535 and colSXOO3762/colRXOO3763, have been annotated (Lee et al., 2005). Based on literature and homologous gene search, the three colS/colR genes were identified to be well conserved in Xcc 8004 (a black rot pathogen of cabbage) and X. axonopodis pv. citri (Xac) 306 (a citrus canker pathogen of citrus) (Table 1). Nucleotide identity between the colS/colR genes of Xoo KACC10331 and Xac 306 or Xcc 8004 were all more than 90%. E-value obtained by BlastN between each colS/colR genes of Xoo KACC10331 and corresponding genes in either Xac 306 or Xcc 8004 were all 0.0. These indicate that nucleotide sequences of the three colS/colR genes are well conserved in the three pathogens.
The colSXC_1050/colRXC_1049, colSXAC3249/colRXAC3250 and colSXOO1207/colRXOO1208 are known to be all required for virulence in each pathogen. Zhang et al. (2008) showed that colSXC_1050/colRXC_1049 was involved in virulence, hypersensitive response and tolerance to various stresses in Xcc 8004. The colSXC_1050/colRXC_1049 positively regulated expression of hrpC and hrpE operons and expression of colSXC_1050/colRXC_1049 were not controlled by the key hrp regulators HrpG and HrpX. Yan and Wang (2011) showed that colSXAC3249/colRXAC3250 were critical for X. citri subsp. citri (Xac) in virulence, growth in planta, biofilm formation, catalase activity, LPS production, and resistance to environmental stress. In Xoo, colSXOO1207/colRXOO1208 were required for virulence and hypersensitive response on non-host plant (Subramoni et al., 2012).
Rpf Mutants, complementation and qRT-PCR
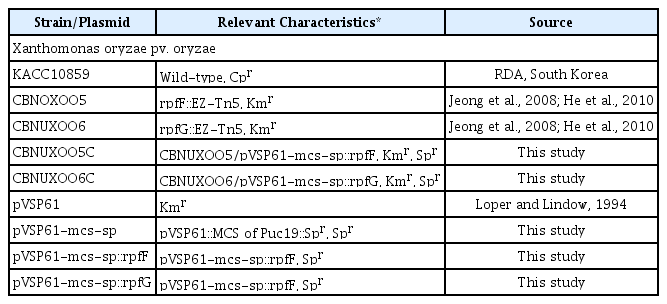
Wild type strain, Xoo KACC10859, two Rpf mutant strains, CBNUXO05 (rpfF::EZ-Tn5), CBNUXO06 (rpfG::EZ-Tn5) and complement strains, CBNUXO05C (rpfF::EZ-Tn5/pVSP61-mcs-sp::rpfF), CBNUXO06C (rpfG::EZ-Tn5/pVSP61-mcs-sp::rpfG) were used in this study (Table 2). Mutants of rpf genes and their biological characteristics including DSF production were published previously (He et al., 2010; Jeong et al., 2008). To complement the mutants, cloning vector, pVSP61-mcs-sp was constructed by modification of pVSP61, which is very stable plasmid vector in Pseudomonads (Loper and Lindow, 1994). Polylinker of pUC9 in pVSP61 was replaced with polylinker of pUC19 at EcoRI – HindIII sites to increase availability of cloning sites (pVSP61-mcs). Spectinomycin resistance gene was inserted into BglII site in KmR gene of the pVSP61-mcs resulting in Kanamycin resistance inactivated and Spectinomycin resistance (pVSP61-mcs-sp).
For complementation of CBNUXO05 (rpfF::EZ-Tn5), DNA fragments containing rpfF including 200 bp of its 5′ region were cloned into HindIII-KpnI sites with PCR products amplified with primers, rpfF-kpn1-F: 5′ AGTGGTACCACATCAGCCGGCGTCAAGC 3′ and rpfF-hind3-R: 5′ AGTAAGCTTCCGTGAATGCGGGACGCG 3′ (pVSP61-mcs-sp::rpfF). For complementation of CBNUXO06 (rpfG::EZ-Tn5), DNA fragments containing rpfG including 386 bp of its 5′ region were cloned into HindIII-BamHI sites with PCR products amplified with primers, rpfG-hind3-F: 5′ AGTAAGCTTAAGGACGGCGGTGACGACG 3′ and rpfG-bamh1-R: 5′AGTGGATCCATCACGCAGCTGACCAGGCG 3′ (pVSP61-mcs-sp::rpfG). Plasmid pVSP61-mcs-sp::rpfF and pVSP61-mcs-sp::rpfG were transformed into CBNUXO05 (rpfF::EZ-Tn5) and CBNUXO06 (rpfG::EZ-Tn5) with standard electroporation protocol. Mutants and its complementation strains were confirmed by PCR genotyping (supplementary Fig. 1).
RNA was isolated from the bacterial cells cultured in hrp-inducing culture conditions (Seo et al., 2008). Wild type, mutant and complement strains were cultured in PS broth (peptone 10 g, sucrose 10 g, l-glutamic acid 1 g per 1 L, pH 7.0) to OD600=0.2 and the bacterial cells were washed twice with sterilized water and transferred to Xom2 medium (0.18% xylose, 670 uM l-methionine, 10 mM L-glutamic acid, 14.7 mM potassium phosphate (monobasic), 40 μM manganese sulfate, 240 μM Fe(III) EDTA, 5 mM magnesium chloride per 1 L, pH 6.5). After 18 h further culture, bacterial cells were harvested for RNA isolation. Total RNA of each strains was isolated using the RNeasy® Mini kit (Qiagen, Valencia, CA), and residual genomic DNA was removed using the RNase-Free DNase Set (Qiagen, Valencia, CA) and RNeasy® MinEluteTM Cleanup kit (Qiagen, Valencia, CA), according to the manufacturer’s instructions. DNase I-treated total RNA was measured with a Nano Drop 2000 (Thermo scientific, Wilmington, USA) and 1 μg total RNA was used to synthesize cDNA. cDNA was generated using SuperScript® III First-Strand Synthesis System for RT-PCR (Invitrogen, Carlsbad, CA), following the manufacturer’s protocol. After the reaction, samples were diluted with 190 μL of distilled water and used as a template cDNA for qRT-PCR. qRT-PCR experiments were performed by Rotor-gene Q (Qiagen, Valencia, CA) and using 2× Rotor-geneTM SYBR® Green PCR kit (Qiagen, Valencia, CA) containing 1 μL of diluted cDNA. The constitutively expressed rrsA gene coding 16S rRNA was used as an internal control for relative quantification (Subramoni et al., 2012). Gene-specific primers designed and their amplification efficiency was checked with RNA isolated from the wild type strain. Primers with R2>0.99 and slope 3.2–3.6 were used (Table 3). qRT-PCR condition included initial heating for 10 min at 95°C, followed by 40 cycles of PCR (95°C, 15 s; 58°C, 30 s; 72°C 30 s). To analyze obtained results, ΔΔCt method was used and quantification results were calculated.
Expression of rpfF and rpfG genes in its mutant and complement strains
Expression of rpfF and rpfG in its mutants and complement strains was checked to confirm the mutation and complementation works. The rpfF and rpfG was expressed about 11 fold and 100 fold less in its mutant strain, CBNUXO05 (rpfF::EZ-Tn5) and CBNUXO06 (rpfG::EZ-Tn5) than in the wild type strain respectively (Table 4). Expression of rpfF in the complement strain, CBNUXO05C (rpfF::EZ-Tn5/pVSP61-mcs-sp::rpfF) was similar to the wild type strain while rpfG was expressed in the complemented strain, CBNUXO06C (rpfG::EZ-Tn5/pVSP61-mcs-sp::rpfG), was about 4 fold higher than in the wild type (Table 4). Forward and reverse primer of rpfF- and rpfG-specific primers were designed from outside of both end of EZ-Tn5 insertion site. Since EZ-Tn5 contains transcription terminator, expression level of the two genes must be near 0 in the mutant strains. When the PCR products were checked by gel electrophoresis after qRT-PCR, no proper-size band or very faint band were appeared in triplicate lanes. Although results of qRT-PCR showed the low expression level of rpfF and rpfG in its mutant strains, these results indicate that RpfF and RpfG were non-functional in the respective mutant while those in the complemented strains were similar to the wild-type strain.
Expression of colS/colR genes was increased in rpfF and rpfG mutants
Expressions of colSXOO1207/colRXOO1208 about 2 fold increased in CBNUXO05 (rpfF::EZ-Tn5), while expression of both genes was not different in CBNUXO06 (rpfG::EZ-Tn5) (Table 4). Although we do not know biological significant of 2 fold increase of this two-component system in the rpfF mutant yet, qRT-PCR results suggest that expression colSXOO1207/colRXOO1208 is influenced directly by DSF rather than signal through RpfG. In Rpf virulence regulation system, RpfF are responsible for the production of DSF (Barber et al., 1997; Wang et al., 2004) and RpfG transfers signal from RpfC that sense the DSF signal to its downstream (He et al., 2007; Ryan et al., 2007; Slater et al., 2000).
Expressions of colSXOO3534 (raxH)/colRXOO3535 (raxR) highly increased in both rpfF and rpfG mutants (Table 4). Coding sensor gene, colSXOO3534 was expressed about 7 and 4 fold higher in the rpfF mutant and the rpfG mutant, respectively, while its cognate coding regulator gene, colRXOO3535, was expressed about 3 and 2 fold higher than in the wild type. These results suggest that DSF and signal from DSF through RpfG regulate negatively the expression of colSXOO3534/colRXOO3535 in the wild strain. Since colSXOO3534/colRXOO3535 has been proved to control of avrXa21 (Lee et al., 2008; Burdman et al., 2004), DSF may suppress avirulence activity by suppression of the expression of two genes for promoting virulence.
Expression of colSXOO3762/colRXOO3763 was increased about 13 and 3 folds, respectively in CBNUXO05 (rpfF::EZ-Tn5) and expression of colSXOO3762 was also increased about 2 folds in CBNUXO06 (rpfG::EZ-Tn5) comparing to wild type strain and expression of its cognate regulator, colRXOO3763, was not changed significantly in CBNUXO06 (rpfG::EZ-Tn5). These results suggest DSF regulates negatively this two-component system in the wild type. Biological function of positive regulation of colSXOO3762 is not clear, since function of colSXOO3762/colRXOO3763 is not known.
In this study, expression of three two-component system genes, two of them are known to control virulence and avirulence, increased significantly in the rpfF mutant, CBNUXO05 (rpfF::EZ-Tn5), and expression of colSXOO3534 (raxH)/colRXOO3535 (raxR) and colSXOO3762 also increased in the rpfG mutant, CBNUXO06 (rpfG::EZ-Tn5). Overall these results indicate DSF and downstream of DSF signal regulate negatively three colS/colR genes in Xoo. Although biological function of these regulations by DSF is unclear yet and further detail work on this area is needed, we think that these regulations may be a part of hierarchal control of pathogenicity, which is needed for pathogen to successfully cause a disease on host plant.
Supplementary Figure
Expression of colSR Genes Increased in the rpf Mutants of Xanthomonas oryzae pv. oryzae KACC10859.
Acknowledgement
This work was supported by a research grant from Chungbuk National University and Pusan National University Research Grant in 2011.