Lysinabacillus fusiformis and Paenibacillus alvei Obtained from the Internal of Nasutitermes Termites Revealed Their Ability as Antagonist of Plant Pathogenic Fungi
Article information
Abstract
This study was performed to reveal phenotypic characters and identity of symbiont bacteria of Nasutitermes as well as investigate their potential as antagonist of plant pathogenic fungi. Isolation of the symbiont bacteria was carried out from inside the heads and the bodies of soldier and worker termite which were collected from 3 locations of nests. Identification was performed using phenotypic test and sequence of 16S ribosomal DNA (16S rDNA). Antagonistic capability was investigated in the laboratory against 3 phytopathogenic fungi i.e., Phytophthora capsici, Ganoderma boninense, and Rigidoporus microporus. Totally, 39 bacterial isolates were obtained from inside the heads and the bodies of Nasutitermes. All the isolates showed capability to inhibit growth of P. capsici, however, 34 isolates showed capability to inhibit growth of G. boninense and 32 isolates showed capability to inhibit growth of R. microporus. Two bacterial strains (IK3.1P and 1B1.2P) which showed the highest percentage of inhibition were further identified based on their sequence of 16S rDNA. The result showed that 1K3.1P strain was placed in the group of type strain and reference strains of Lysinibacillus fusiformis meanwhile 1B1.2P strain was grouped within type strain and reference strains Paenibacillus alvei. The result of this study supply valuable information on the role of symbiont bacteria of Nasutitermes, which may support the development of the control method of the three above-mentioned phytopathogenic fungi.
Nasutitermitinae is one of the members of higher termites comprising 650 species which is also known as a type of invasive termite (Evans et al., 2013). This group of termites has been widely recognized as pests of wood and woody plants causing severe damage worldwide (Ahmad et al., 2021; Evans et al., 2013). Alongside its negative role, diverse microbes reside within termites have been isolated and confirmed to have beneficial task that can give advantages to various fields in human life, including in agriculture. Some of termite-symbiont bacterial genus have been confirmed to have capability to degrade lignin, lignocellulose and the other aromatics compounds as well as nitrogen fixer, such as Bacteroides, Clostridia (Trakulnaleamsai et al., 2004), Paenibacillus (Butera et al., 2016), and Spirochaetes (Husseneder, 2010).
So far, there is no report yet on the characteristics and role of symbiont bacteria of termites, especially Nasutitermes as antagonists of plant pathogenic fungi. This research was performed to reveal phenotypic characters and identity of the symbiont bacteria of Nasutitermes as well as their role as antagonist against three kinds of plant pathogenic fungi i.e., Ganoderma boninense (pathogen of basal stem rot of oil palm), Phytophthora capsici (pathogen of foot rot of pepper), and Rigidoporus microporus (pathogen of white root rot of rubber). Oil palm, pepper, and rubber are three estates crops whose products have high economical value supporting foreign exchange earnings of some countries, including in Indonesia. The three above-mentioned plant pathogenic fungi become the main obstacle in the effort to increase production of each plant.
Biological controls using antagonists become promising methods to control plant pathogens. Beside its nature which always provides protection as long as the antagonist is alive, the use of antagonist is an eco-friendly controlling method that will lead to a sustainable agricultural condition. The antagonist that is selected from symbiont bacteria of termites will have more advantages. Because these bacteria have capability to degrade lignin or lignocellulose (Evans et al., 2013), they will be able to have more inhibition capability against plant pathogenic fungi than is obtained from soil or rhizosphere.
Materials and Methods
Sample collection and isolation of the symbiont bacteria of termite
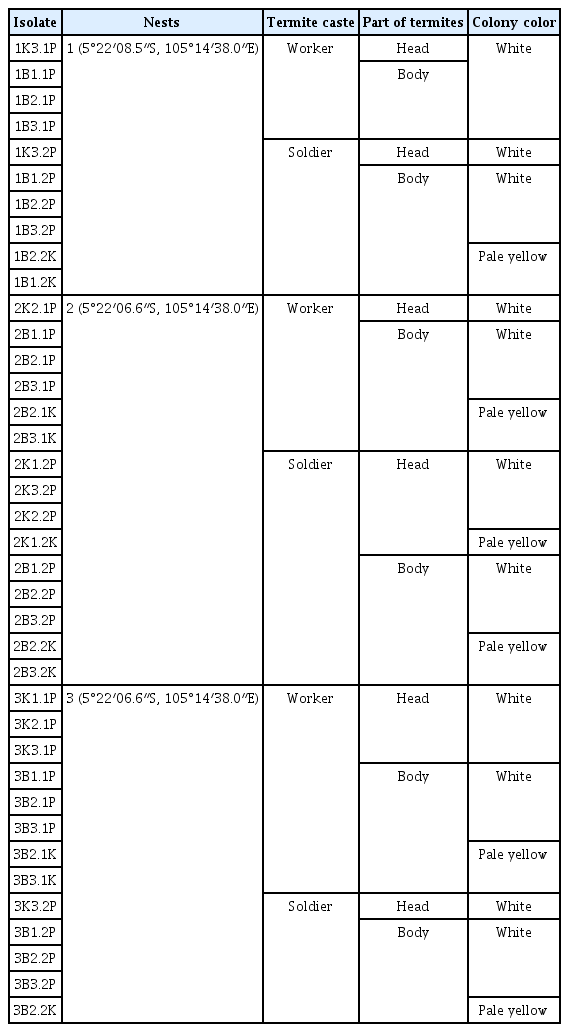
Totally, 6 termites (3 soldiers and 3 workers termites) were collected from 3 locations of nests located in Bandar Lampung, Lampung Province, Indonesia (Table 1). The locations of the nests were determined by conducting a survey on dry fields cultivated by wood or woody plants. The collected termites were surface sterilized by dipping into sterile distillate water then 0.5% of natriun hypochlorite (NaOCl) then sterile distillate water, one minute of each and air dried. This step was repeated 3 times. The head and the body (thorax and abdomen) were separated and each of the part was put into 1.5 ml tube contained 20 μl phosphate buffer saline (4 g NaCl, 0.1 g KCl, 0.12 g KH2HPO4, 0.72 g Na2HPO4, 500 ml distillate water, pH 7.2) (Schaad et al., 2001) and completely crushed using a sterile tweezer. Afterward, 20 μl of the suspension was spread onto yeast peptone agar medium (5 g yeast extract, 10 g peptone, 20 g agar, 1,000 ml distilled water) (Suharjo et al., 2014) using drigalski spatula. The growing colonies were transferred onto potato peptone glucose agar (PPGA) medium (200 g potato, 5 g peptone, 5 g glucose, 3 g Na2HPO4·2H2O, 3 g NaCl, 0.5 g KH2PO4, 20 g agar, 1,000 ml distilled water) (Nishiyama, 1978). Before it being used, the bacteria were preserved in skim milk medium (5 g skim milk, 0.75 g Na glutamate, 50 ml distilled water) (Suharjo et al., 2014) and kept in −40°C.
Phenotypic properties test
Four phenotypic properties tests were used to characterize the obtained bacteria i.e., gram reaction test, oxidation and fermentation (OF) test, potato soft rot test and hypersensitive reaction test on tobacco leaves. The gram reaction test was performed by a non-staining method using 3% KOH (Ryu, 1940). The OF test was assessed using the OF medium described by Hugh and Leifson (1953). The potato soft rot test was conducted using a method of Lelliot et al. (1966). The hypersensitive reaction test was conducted by a method described by Schaad et al. (2001).
Hypovirulence determination
The pathogenicity of the obtained bacterial isolates was investigated using water agar (WA) method (Ichielevich-Auster et al., 1985; Juan-abgona et al., 1996) with slight modification. Cucumber (Cucumis sativus) cv. TAFI (PT Dynasti Inti Agrosarana, Kabupaten Bantul, Indonesia) was used as a test crop. For surface disinfection, the cucumber seeds were deep into warm water (45°C) for 30 min followed by deeping in 70% ethanol for 15 min and soaking in 0.5% sodium hypochlorite (NaOCl) for 30 s. The seed then rinsed 3 times using sterile distillate water and pre-germinated by placing on the plastic tray (40 × 40 × 60 cm) which was bottom-covered with moisturized parchment paper (by spraying 30 ml distillate water using hand sprayer (7.5 cm × 5.5 cm × 24.5 cm) and wrapped using cling wrap and incubated at room temperature (dark-light condition) for 1 day. After incubation, 4 seeds were placed into a sterile petridish (10 cm of diameter) containing 10 ml WA medium (20 g agar, 1,000 ml distillate water) and incubated at room temperature (dark-light condition) for 1 day. In total, 10 μl bacterial suspension (108 colony forming unit [cfu]/ml; which was made from 24-hour-old bacteria grown on PPGA slant medium suspended in sterile distillate water) was dropped on the hypocotyl of the cucumber seedling which was grown on WA medium. The disease severity index (DSI) was calculated by dividing the total of individual seedling score with the total of individual seedling observed following scoring scheme described by Cardoso and Echandi (1987) with slight modification, where 0 = healthy, without any necrotic symptoms, 1 = one or two necrotic symptom with light brown color, ≤0.25 cm, 2 = necrotic symptom with light brown color, 0.25–0.5 cm, with water soaked area was ≤10% of hypocotyl, 3 = necrotic symptom with light to dark brown color, fused, ≥1 cm, with water soaked area was 10–100% of hypocotyl, 4 = necrotic symptom with black color, hypocotyl collapse and the seedling is death.
Antagonist test
Antagonist test was performed against 3 phytopathogenic fungi i.e., P. capsici, G. boninense, and R. microporus which were the main pathogen of 3 main estate crops (pepper, oil palm, and rubber) in Indonesia. The symbiont bacteria were streaked vertically at 2.5 cm from the left and the right edge of the petri dish (10 cm of diameter) (a petri dish contained 2 strokes of the symbiont bacteria) contained 10 ml of potato dextrose agar (PDA; 200 g/l potatoes, infusion; 20 g/l dextrose [glucose]; 15 g/l agar; final pH [at 25°C] 5.6 ± 0.2) (Himedia, Thane West, India). A mycelial plug (0.5 cm of diameter) of each of 5-day-old phytopathogenic fungi was placed in the middle of the petri dish (10 cm of diameter).
As control, a mycelial plug (0.5 cm of diameter) was put in the petri dish (10 cm of diameter) containing PDA medium (Himedia) without any symbiont bacteria. Observation was performed every day for 7 days on the colony diameter of phytopathogenic fungi. Percentage of inhibition was calculated using formula [(D1 − D2)/D1] × 100%, where D1 = colony diameter of phytopathogenic fungi in the control, D2 = colony diameter of phytopathogenic fungi in the petri dish contained symbiont bacteria.
Molecular identification
DNA extraction
A full loop of 24-hour-old bacterial isolates which was cultivated on PPGA slant was transferred into 1.5 ml containing 20 μl 1× Tris HCl EDTA (TE) buffer pH 8.0 (1st BASE, Selangor, Malaysia) and robustly homogenized. As much as 10 ml 10% sodium dodecyl sulfate and 3 ml proteinase K (10 mg/ml) were added, gently homogenized and incubated in a dry block incubator (MBDB-01, Cleaver Scientific, Warwickshire, UK) at 37°C for 1 h. After incubation, a 100 μl 5 M NaCl was directly added and homogenized by hand shaking. A 80 μl CTAB 2% was added and incubated at 65°C for 15 min in a dry block incubator (MBDB-01, Cleaver Scientific). After incubation, 720 μl chloroform:isoamyl alcohol (24:1) was added and homogenized by hand shaking and centrifuged at 14,000 rpm for 5 min. The supernatant was removed into a new 1.5 ml tube and phenol:chloroform:isoamyl alcohol (25:24:1) was added at the same volume as the volume of supernatant. The mixture was hardly homogenized using hand shaking and centrifuged at 14,000 rpm for 5 min. The supernatant was transferred into a new 1.5 mL tube and cold isopropanol was added (60% of the volume of supernatant), gently homogenized by hand shaking and incubated for 20 min at freezer (−40°C). After incubation, the tube was centrifuged at 14,000 rpm for 15 min and the supernatant was expelled. The tube was added by 400 μl of 70% cold ethanol and centrifuged at 14,000 rpm for 5 min. The supernatant was discharged and air dried overnight at room temperature. After incubation, 20 μl 1× buffer Tris HCl EDTA (TE) pH 8.0 (1st BASE) was added. All centrifugation processes were performed using a centrifuge Microspin12 (Biosan, Riga, Latvia).
Polymerase chain reaction amplification of 16S ribosomal DNA
Amplification of 16S ribosomal DNA (16S rDNA) was conducted using Sensoquest Thermal Cycler machine (Gottingen, Germany) using primer fD1 (5′-CCGAATTCGTCGACAACAGAGTTTGATCCTGGCTCAG-3′) and rP2 (5′-CCCGGGATCCAAGCTTACGGCTACCTTGTTACGACTT-3′) (Weisburg et al., 1991). Polymerase chain reaction (PCR) was carried out in total volume 25 μl involving 12.5 μl Master Mix (Red Mix) (Bioline, Nottingham, UK), 1 μl of 10 μM of each primer, 1 μl DNA template (~1 μg/μl), and 9.5 μl sterile distilled water. DNA amplification was performed covering one cycle of the initiation at 95°C for 5 min, 30 cycles of denaturation at 95°C for 1 min, primer annealing at 58°C for 1 min, primer extension at 72°C for 1 min, and one cycle of elongation at 72°C for 5 min. The results of PCR were electrophoresed using 0.5% agarose gel in 20 ml 1× buffer Tris-boric acid-EDTA (TBE) pH 8.0 (1st BASE) supplemented with 1 μl ethidium bromide (EtBr 10 mg/ml). The electrophoresis was conducted using a 1× TBE buffer (1st BASE) at 50 V for 70 min. The results were visualized using DigiDoc UV transilluminator (Major Science, Saratoga, CA, USA).
Sequencing and its result analysis
The PCR product was sent to 1st BASE for the sequencing process. The sequencing result was analyzed using bioedit version 7.2.6 for windows (Hall, 1999). Phylogenetic tree was developed using Mega 7 for windows (Kumar et al., 2016).
Data analyses
Antagonistic assessment was arranged using Completely Randomized Design (CRD) with 4 replicates. The data was investigated using one-way analysis of variance (ANOVA). If there is a significant difference among the means of two or more bacterial strains, advance analysis was performed by least significant difference assay. Statistical analysis was carried out using R Statistical Software (version 4.1.1, R Foundation for Statistical Computing, Vienna, Austria).
Results
Putative symbiont bacteria obtained in this study
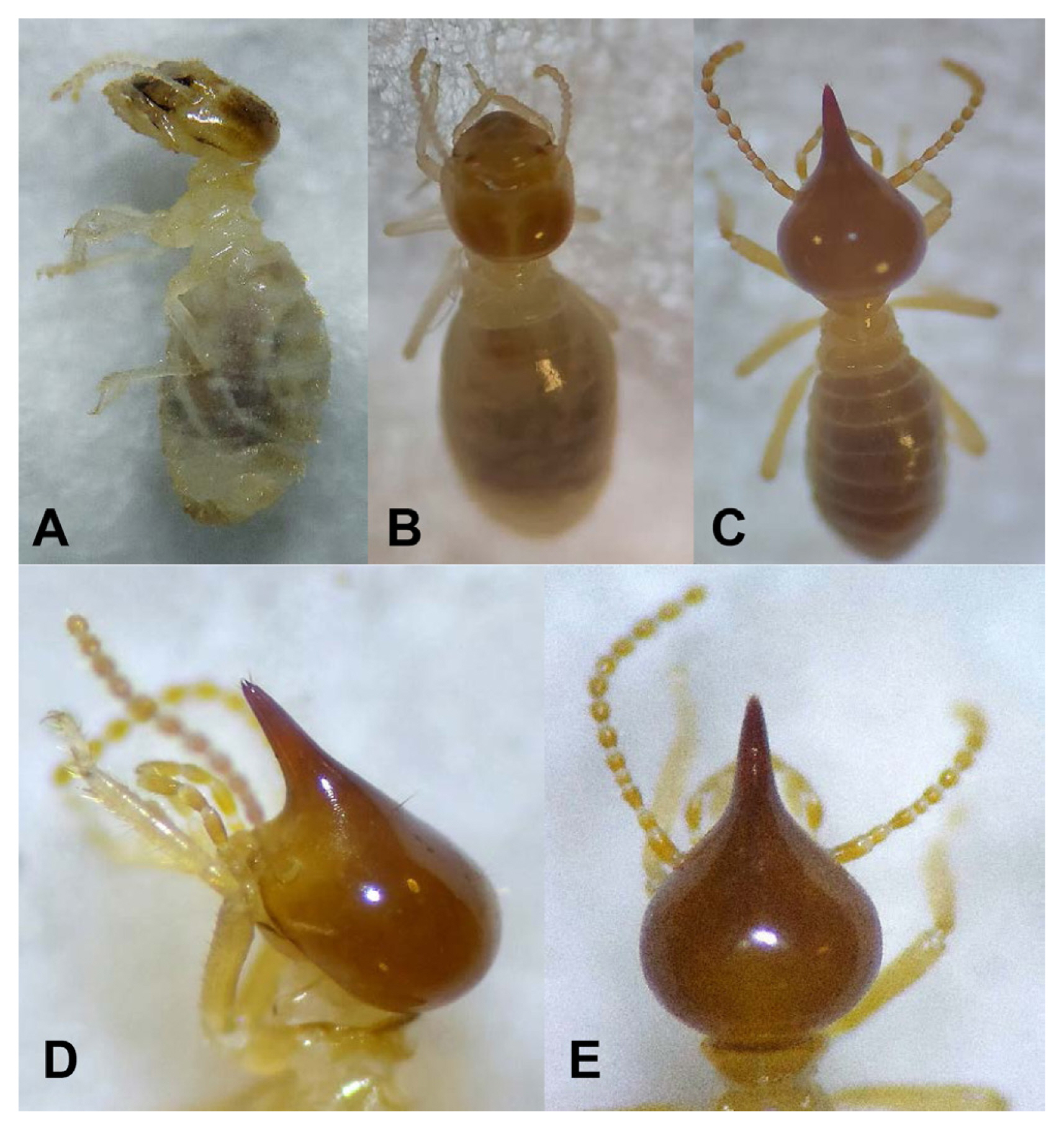
Two kinds of termites castes used as isolation objects i.e. soldier and worker (Fig. 1). Totally, 39 bacterial isolates were collected as representative strains of the bacteria which emerged (Table 1). A representative isolate was selected from the growing bacterial colonies which showed similarity on the colony shape and colony color. Ten bacterial isolates were obtained from inside the head and 29 isolates were isolated from inside the body.
Phenotypic properties
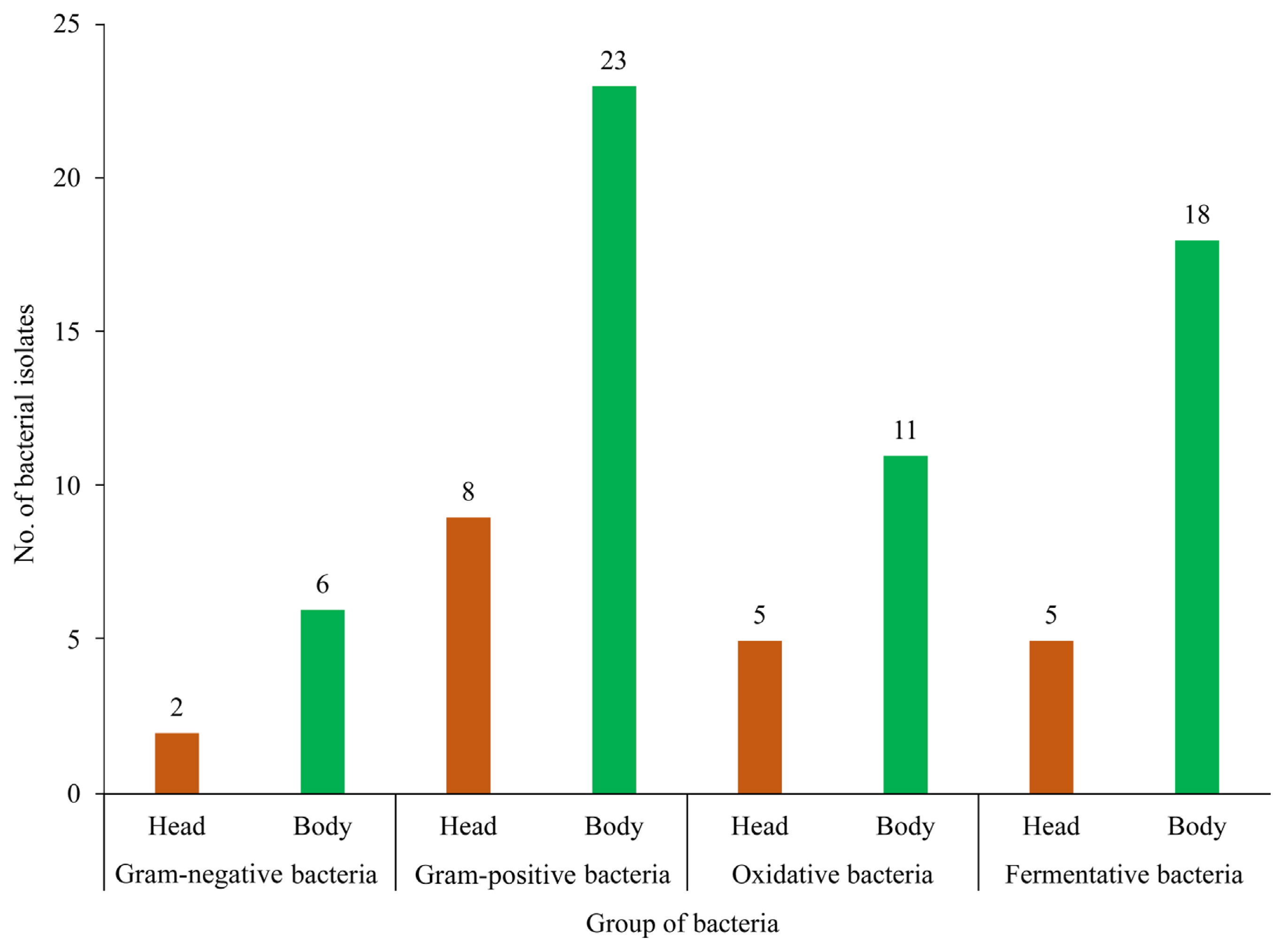
Eight strains out of 39 isolates were in the group of gram-negative bacteria (2 isolates from the head and 6 isolates from the body), while the other 31 isolates were in the group of gram-positive bacteria (8 isolates from the head and 23 isolates from the body). Sixteen isolates out of 39 isolates were oxidative bacteria (5 isolates from the head and 11 isolates from the body), meanwhile the other 23 isolates were fermentative bacteria (5 isolates from the head and 18 isolates from the body) (Fig. 2). All the isolates showed negative results in the soft rot and hypersensitive reaction test.
Hypovirulent test
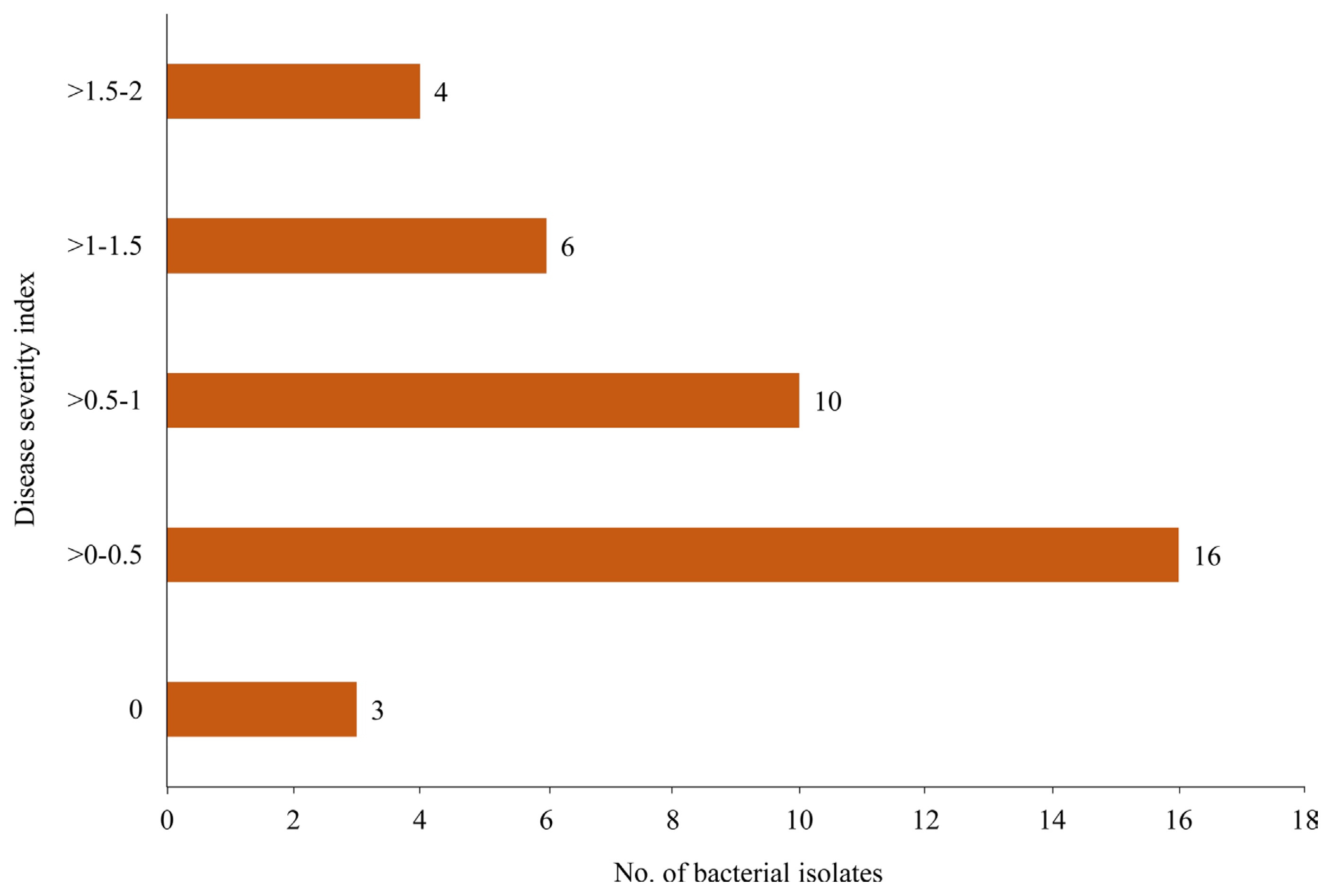
All the bacterial isolates were hypovirulent. The DSI observed on the cucumber sprout as the indicator plant was less than 2.0, which was in the range of 0–1.7 (Fig. 3). Most of the bacteria (16 isolates) produced DSI in the range of 0–0.5, followed by >0.5–1 (10 isolates), >1–1.5 (6 isolates), >1.5–2 (4 isolates), and 0 of DSI which was produced only by 3 isolates of bacteria (Fig. 4).

Representative hypovirulent test on cucumber sprout at 14 days after bacterial inoculation. (A) 1B1.1P isolate (disease severity index [DSI] = 1.3). (B) 1B2.1P isolate (DSI = 0.2). (C) 2K3.2P (DSI = 0). Necrotic symptom (indicated by arrow) is designated by the presence of rotten area on the sprout in either the leaves or stems.
Antagonist test
Thirty-four isolates showed capability to inhibit growth of G. boninense but not for the other 5 isolates. Nine isolates out of 34 isolates produced more than 50% of percentage of inhibition and the other 25 isolates showed less than 50% of percentage of inhibition against G. boninense. Thirty-two bacterial isolates had antagonistic capability against R. microporus, but not for the other 7 isolates. All the 32 isolates produced less than 50% of the percentage of inhibition against R. microporus. All the isolates showed antagonist capability against P. capsici. Eleven isolates showed more than 50% of percentage of inhibition, while the other 28 isolates produced less than 50% of percentage of inhibition against P. capsici (Fig. 5). Inhibition capability here indicated by the presence of percentage inhibition though it was in low inhibition level. No inhibition capability was implied by zero percentage of inhibition (Fig. 6).
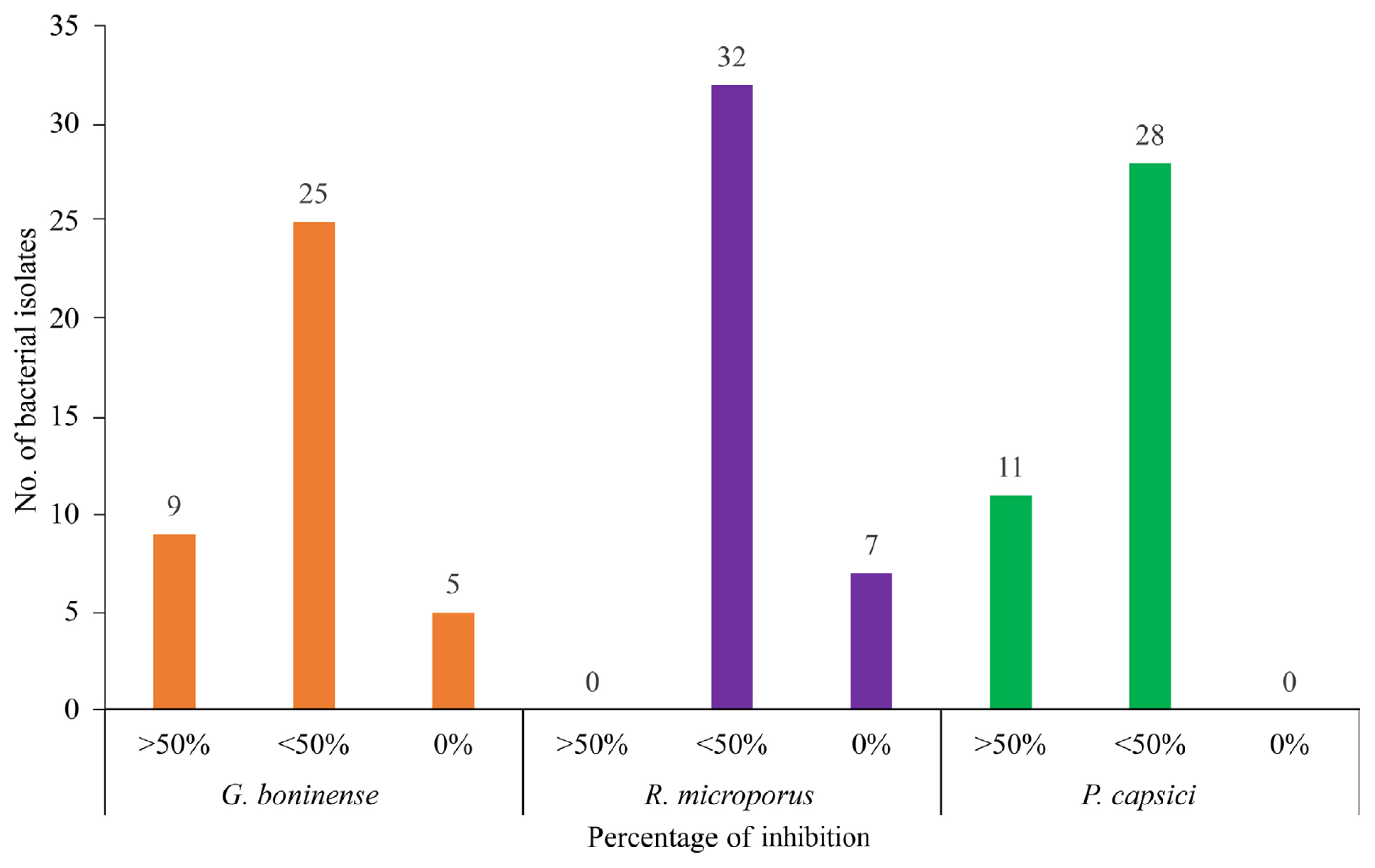
Percentage of inhibition resulted by symbiont bacteria. Generally, the bacterial isolates showed capability to inhibit Ganoderma boninense, Rigidoporus microporus, and Phytophthora capsici. Most of the bacterial isolates were found to have low inhibition (indicated with <50% of inhibition) against tested plant pathogenic fungi. High inhibition (indicated with >50% of inhibition) was observed on small number of bacteria against G. boninense and P. capsici but not against R. microporus. Some of the bacterial isolates have no inhibition capability (indicated with 0 % of inhibition) against G. boninense and R. microporus.

In vitro antagonist test at 7 days after inoculation. (A) 3B1.2K strain against Rigidoporus microporus (0% of inhibition). (B) 1B2.1P strain against Phytophthora capsici (23.71% of inhibition). (C) IBI.2P strain against Ganoderma boninense (93.38% of inhibition).
The highest inhibition capability against G. boninense was shown by strain of 1B1.2P (93.38%). The highest capability to inhibit growth of P. capsici was emitted by 1K3.1P strain (72.59%). The highest inhibition capability against R. microporus was generated by 2K2.2P isolate (19.06%). One bacterial isolate showed different antagonist capability against different plant pathogenic fungi (Fig. 7).
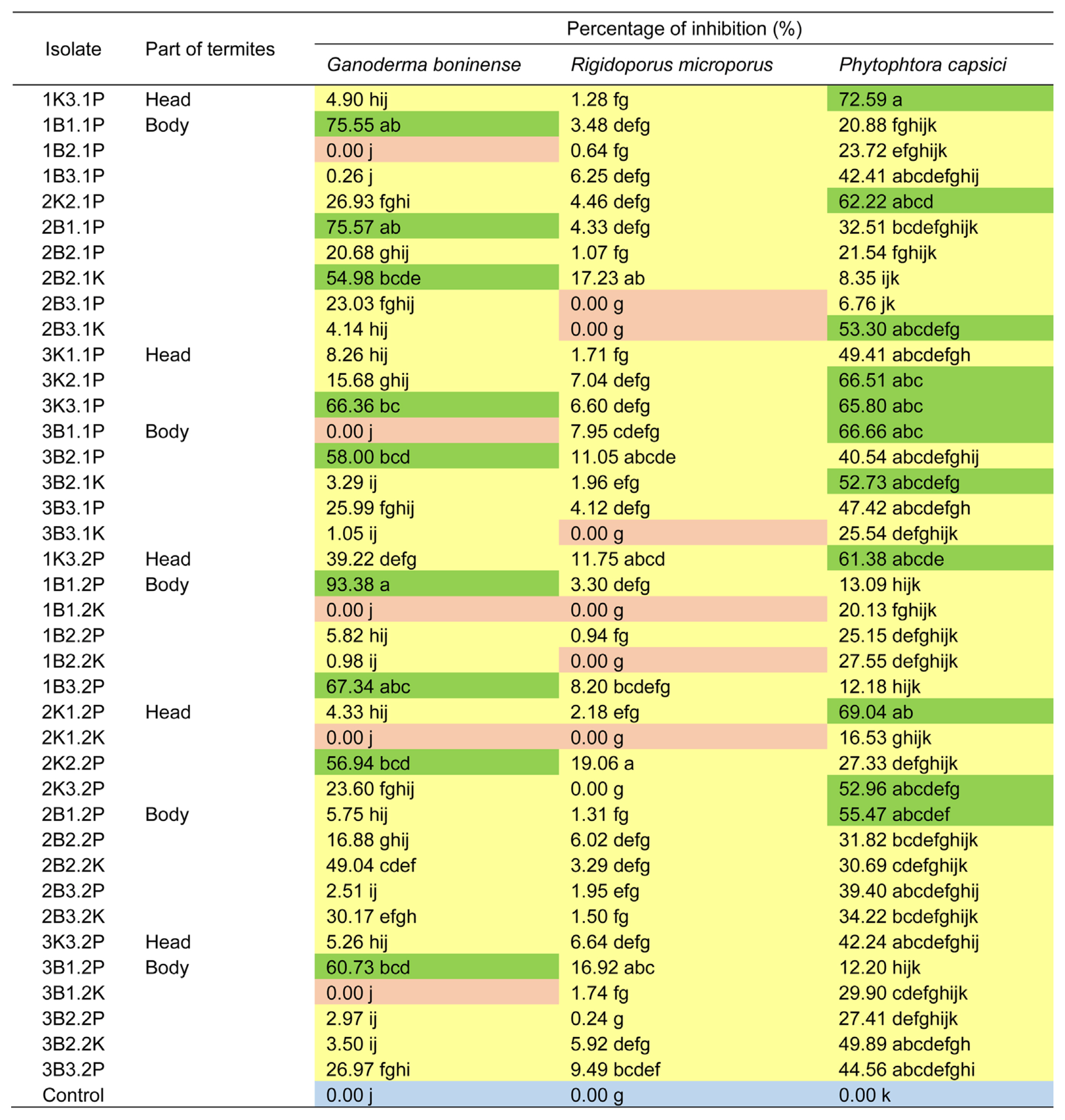
The result of antagonist test. Green color indicates isolates with percentage of inhibition >50%, yellow color is the isolates with percentage of inhibition <50%, orange color is isolates with no inhibition capability (0% of percentage of inhibition). Blue color indicates control. Nine isolates showed capability to inhibit Ganoderma boninense >50% (7 isolates obtained from inside the body, 2 isolates from inside the head). There were no isolates that showed capability to inhibit Rigidoporus microporus with percentage of inhibition >50%. Eleven isolates (4 isolates from inside the body, 7 isolates from inside the head). had capability to inhibit Phytophthora capsici with percentage of inhibition >50%. Values followed by the same letter within the same column are not significantly different based on the least significant difference test at 5% of significant level.
Molecular identification
Two bacterial strains resulting in more than 50% of percentage of inhibition showing the highest inhibition of the tested plant pathogenic fungi were selected for molecular identification. The strains were 1K3.1P (72.59% of percentage of inhibition against P. capsici) and 1B1.2P (93.38% of percentage of inhibition against G. boninense). Because the percentage of inhibition of R. microporus was less than 50%, here, no bacterial isolate was selected from the antagonist test against R. microporus.
The result of the BLAST search tool revealed that 1K3.1P shared 100% of similarity with Paenibacillus alvei NBRC 3343.58 (accession no. MN543811.1), meanwhile 1B1.2P showed 98.76% of similarity with Lysinibacillus fusiformis strain NBRC 15717 (accession no. MN543848.1). Phylogenetic tree analysis showed that IBI.2P strain clustered within type strain (NBRC 3343, accession no. AB680069) and reference strains of P. alvei strain AUG6 (accession no. AB377108.1), strain AN5 (accession no. JQ868768.1) strain IARIDB10 (accession no. KT934806). Meanwhile, the strain of IK3.1P was in group with the type strain (NBRC 15717; accession no. MN543848.1) and reference strains of L. fusiformis strain LMR 802 (accession no. MW559723), strain R53638 (accession no. LT631750.1), and strain A8 (accession no. LT547806.1) (Fig. 8).
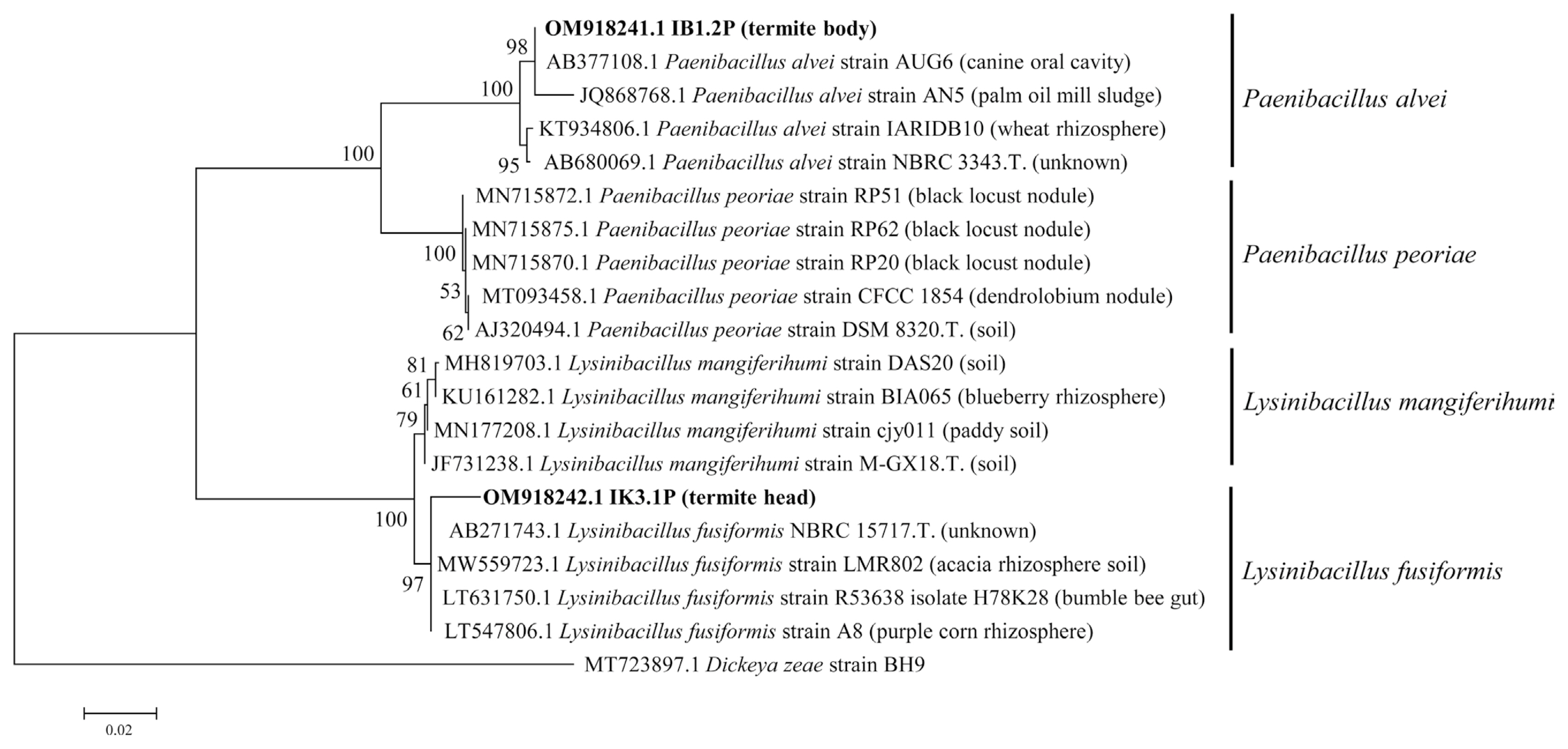
Phylogenetic tree was developed based on a sequence of 16S rDNA using the Maximum Likelihood method (Tamura-Nei model) with 1,000× bootstrap. The strain of IK3.1P was placed within the group of Lysinibacillus fusiformis, meanwhile the strain of IB1.2P was clustered in the group of Paenibacillus alvei. OM918241 and OM918242 are NCBI accession number of IB1.2P and IK3.1P. T., type strain.
Discussion
A numerous kind of culturable microbes has been widely reported to exist and live in insects (Gurung et al., 2019). These insect-associated microbes have been reported as an important source of beneficial microbes that can be widely applied for human advantages in various fields (Berasategui et al., 2016). Some of the insect-associated microbes have been reported to have capacity as antifungal (Cardoza et al., 2006) and antibacterial (Muniaraj et al., 2008; Skowronek et al., 2020). In this study, we investigated diversity of Nasutitermes termites’ symbiont bacteria as well as their potential as antagonist of plant pathogenic fungi i.e., G. boninense, R. microporus, and P. capsici.
Thirty-nine bacterial isolates were obtained from inside the head and the body of two castes of Nasutitermes termites i.e., worker and soldier. Here, a mutually beneficial relationship (mutualism symbiosis) between the bacteria and termites occurred. Termites provide shelter and living habitat (anaerobic condition) as well as food for the bacteria, while the bacteria will contribute to degrade lignocellulose in the termite digestive tract (Hyodo et al., 2000).
The bacterial isolates found to be different in their phenotypic characters. Most of the bacterial isolates were gram-positive bacteria (79.49%). The fermentative bacteria were found more (58.97%) than the oxidative bacteria. Bacterial isolates found from the body were more (75.36%) than those were isolated from the head.
The digestive system is the area with more suitable habitat for the bacteria to grow (Brune, 2013). Within the gut area there are diverse activities with a complex mutualistic relationship between the symbiont and their host (Brune, 2013). Gram-positive bacteria were found more (82.05%) than the gram-negative bacteria. Since the gram-positive bacteria has layers of peptidoglycan many times thicker than gram-negative bacteria, they will have capability to survive and develop more than gram-negative bacteria (Silhavy et al., 2010). The more abundant fermentative bacteria arise as the anaerobic condition inside the termite will give more opportunity for the fermentative bacteria to develop more than oxidative bacteria. The most of bacteria found in termite intestines are possibly involved in the fermentation of the solvable products sent into the gut, which are originated either directly from the food which was digested by the digestive enzymes or by the fermentative activity of the intestinal protozoa (Brune, 2013).
All the isolates exposed negative results on soft rot and hypersensitive reaction tests. The result of the hypovirulent test showed that all the strains were hypovirulent (low virulent) bacteria. This results indicates that the 39 isolates were not plant pathogens which will not cause negative impact when applied to the plant. Soft rot test was performed to confirm plant pathogenic bacteria from a group of soft rot Pectobacteriaceae comprising plant pathogenic bacteria derived from genus Dickeya and Pectobacterium (Adeolu et al., 2016; Charkowski, 2018). A group of bacteria which showed positive results on hypersensitive reaction tests on tobacco leaves is plant pathogen, however, some of plant pathogenic bacteria produce negative results. For example Pectobacterium subsp. brasiliensis (Moleleki et al., 2013) which is recently named as P. brasiliense (Portier et al., 2019) and Erwinia mallotivora (Suharjo et al., 2021) revealed positive result on tobacco hypersensitive reaction test, however some of pectobacterium species i.e. P. atrosepticum and P. wasabiae type strain (isolated from wasabi) as well as P. wasabiae isolated from potato which is currently known as P. parmentieri (Khayi et al., 2016) produced negative result (Kim et al., 2009; Moleleki et al., 2013). Low virulent microbes (hypovirulent) (DSI < 2) will not cause a negative effect to the plant which was applied with this bacteria and vice versa (Sneh et al., 2004; Suharjo et al., 2018). Combinations of the three assessments are strongly recommended to determine the nature of the bacteria as pathogen or not.
Antagonist tests showed that each bacterial strain produced different inhibition capability against different plant pathogenic fungi. One bacterial strain revealed different inhibition capability to different plant pathogenic fungi. Some antagonists are inhibiting specific to a species plant pathogenic fungi. As reported by Suharjo et al. (2018), three isolates of endophytic bacteria (GKSKK, AM, and B1) showed inhibition capability against Curvularia and Thielaviopsis but not to Phytophthora. The isolate of GKSKC showed low inhibition capability against Thielaviopsis (4.88%) but high inhibition was observed against Curvularia (66.08%) and Phytophthora (72.48%). Here, investigation and screening on the capability of antagonists to inhibit growth of different plant pathogens is necessary to confirm their inhibition capability against particular plant pathogens.
Molecular identification was performed on two representative strains showing the highest inhibition capability against G. boninense (1B1.2P) and P. capsici (1K3.1P) which was carried out on the basis of 16S rDNA sequence. Phylogenetic tree analysis results showed that 1K3.1P was placed in the group of reference strains of Lysinibacillus fusiformis, meanwhile 1B1.2P strain was grouped within reference strains of P. alvei.
Lysinibacillus fusiformis (=Bacillus fusiformis) is a gram-positive rod-shape bacteria, obligate aerobe, oxidase positive, able to hydrolysis casein and gelatin. This group of bacteria has optimum growth at 17–37°C, pH 6–9.5 and 2–7% of NaCl (Ahmed et al., 2007; De Vos et al., 2009). P. alvei is a motile spore-producing bacteria which swarms on routine culture medium. They give variable results on Gram reaction tests. This bacteria is able to grow both in aerobe and anaerobe conditions (facultative anaerobe), able to grow in the moderate temperature (mesophyll, unable to grow at >45°C) and in moderate salt concentration (unable to grow at 7% NaCl) (Cohen et al., 2000).
L. fusiformis has been reported as endophyte (Sgroy et al., 2009) and plant growth promoter (Jinal et al., 2019; Park et al., 2005; Passera et al., 2020) as well as antagonist of some plant pathogens such as Aspergillus nigri, Botrytis cinerea (Passera et al., 2020), Candidatus liberibacter asiaticus (Trivedi et al., 2011), Fusarium oxysporum, Fusarium solani, Macrophomina phaseolina (Singh et al., 2013), Phomopsis viticola, and Rhizoctonia solani (Passera et al., 2020). P. alvei has also been reported as plant growth promoter on sea purslane (Sesuvium portulacastrum) (Khalaiselvi et al., 2019) and antagonist of several kinds of plant pathogenic fungi such as Alternaria alternate, Aspergilus ochraceus, Aspergilus niger, B. cinerea, F. oxysporum, Penicillium commune, Plasmopara viticola (Atanasova-Pancevska and Kungulovski, 2018), and Verticillium dahliae (Antonopoulos et al., 2008; Gkizi et al., 2021).
The two bacterial strains (1B1.2P and 1K3.1P) selected in this study showed potential to be widely applied for controlling plant pathogenic fungi. As further experiments, field assessment will be conducted to confirm their antagonistic capability.
Acknowledgments
This work was supported by Daftar Isian Pelaksanaan Anggaran (DIPA) Badan Layanan Umum (BLU) University of Lampung through fundamental research grant No. 1604/UN26.21/PN/2021. We thanks to Faculty of Agriculture, University of Lampung for permitting us using research facilities during this study.
Notes
Conflicts of Interest
No potential conflict of interest relevant to this article was reported.