Due to climate change, the temperature has been constantly rising and the intensity and frequency of extreme climate events have been increasing. As a result, the pattern of crop disease occurrence is changing, with, for instance, an augmentation in the incidence of thermophilic diseases such as rice stripe and bacterial leaf blight, as well as sudden epidemics of invasive diseases due to increased trade and rapid globalization, and an expansion of emerging/re-emerging diseases (Fisher et al., 2012; Lee, 2014; Pautasso et al., 2012). The Rural Development Administration of South Korea (RDA) operates the National Crop Pest Management System (NCPMS; https://ncpms.rda.go.kr) to monitor in real time and manage the insect pests and diseases of major crops in the country, thus making policy interventions in the event of a crisis or a pest/disease outbreak (Lee, 2012). Since the intervention through the NCPMS is mostly based on real-time monitoring for each region, it mainly targets diseases that have already started to spread. However, the present disease control system focuses on preventative rather than therapeutic control because the former is more cost-effective than the latter. Although the NCPMS provides weather forecast-based predictions for selected pests and diseases, this has a limitation because the collaborative disease controls require at least a month for the decision-making to schedule and prepare the control activities. Considering these practical discrepancies, it is necessary to introduce a disease prediction system that uses seasonal climate forecasts (SCFs), with a lead time of 1-3 months. If this can be accomplished, it might be possible to take preventive measures in a timely manner, based on the seasonal disease prediction information.
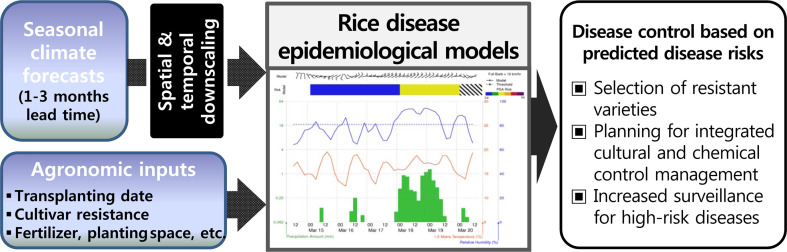
The operational use of SCF information with agricultural models is a pioneering a pioneering attempt that has rarely been implemented in the world, but not yet in South Korea (Kim et al., 2019b; Stone and Meinke, 2005). Linking the SCF with the agricultural models is to establish a system in which valuable climate information can be transformed into agricultural information. The seasonal time scale information from the system enables agricultural stakeholders to make informed decisions, especially for disease control strategies requiring a few-month lead time. To this end, there are two essential components required for the seasonal disease early warning service to be established (Fig. 1). First, SCFs with a coarse spatial and temporal resolution need to be downscaled, resulting in application-ready, daily, and local-specific weather data that can be used by plant disease epidemiological models (referred to as disease model hereafter). To date, many downscaling techniques used to link SCFs with agricultural models have been developed through interdisciplinary research (Han et al., 2017; Hansen and Indeje, 2004; Kim et al., 2019a, 2019b). Next, a disease model compatible with the downscaled SCFs should be developed. When linking the disease model with the SCF, there is a scale problem to be solved. National scale modeling using disease models requires the integration of dispersed data sources in a consistent and spatially and temporally complete dataset to provide necessary model inputs and transfer site-based knowledge to larger areas of a province or country. However, with the increasing size of the area under investigation, input data tend to become more uncertain in relation to the point data from the experimental sites (Parker et al., 2002). Therefore, the model that is applied should also embrace the uncertainty by decreasing the sensitivity to highly uncertain inputs but increasing that to more certain inputs, which can be done through the restructuring or meta-modeling of the original model (Sparks et al., 2011). Once these two components are addressed, the seasonal disease early warning service will be able to generate seasonal risk information for a specific disease. Based on this information, the end users can plan in advance possible measures, including non-chemical and chemical methods, to control the identified high-risk diseases.
As one of the key elements for the establishment of the seasonal rice disease early warning service, we needed a rice disease epidemiological model that is sensitive to seasonal climate variability and structurally generic enough to deal with multiple diseases with one modeling framework. In addition, the model had to already be completely parameterized and validated based on ground-truth data collected from rice paddies in South Korea. Considering all these factors, we found that the EPIRICE model from our previous study closely met most of the abovementioned requirements (Kim et al., 2015). Because of its broad genericity and simplicity but sound infection algorithms, EPIRICE has been adopted in many modeling-based studies worldwide (Duku et al., 2016; Hensawang et al., 2017; Kim and Cho, 2016; Kim et al., 2015; Sittisak et al., 2017). In this respect, our objective was to develop an SCF-compatible disease epidemiological model by extracting and modifying the core infection algorithms of the EPIRICE model. First, a daily risk model for rice blast was developed from the EPIRICE-LB model from Kim et al. (2015). Next, we calibrated and validated the resulting EPIRICE Daily Risk Model using multiple rice blast incidence datasets collected in South Korea.
Materials and Methods
Development of the EPIRICE Daily Risk Model.
In 2013, we introduced the EPIRICE model, originally developed by Savary et al. (2012), to predict the potential future epidemics of rice blast and sheath blight in South Korea under climate change. The EPIRICE model was modified to reflect the local characteristics of rice paddies and climate conditions of South Korea through the process of parameterization and the addition of new modules, resulting in the EPIRICE-LB for rice blast and EPIRICE-SB for sheath blight. In this study, the core infection algorithms of the EPIRICE-LB model were extracted to develop a new daily infection risk model of rice blast, the EPIRICE Daily Risk Model. Specifically, among the main algorithms for determining the overall infection rate, we tried to extract all essential modules related to the weather effects and the ageand cultivar-dependent plant resistance (or susceptibility) effects. However, complex mechanical modules simulating plant growth and disease transmission were avoided, as the daily infection risk model should be simple in structure and sensitive to environmental conditions. The formula for determining the daily infection rate can be expressed as follows:
In Eq. (1), Rc is the daily infection rate, RcT is the temperature effect, RcW is the leaf wetness (estimated from relative humidity and precipitation) effect, RcA is the plant age effect, and RcCS is the plant cultivar susceptibility effect. The effects of temperature, leaf wetness, plant age, and cultivar susceptibility level were translated into daily infection risks using daily weather data for air temperature, precipitation, and relative humidity. In addition, some modules in the EPIRICE model that had an on-and-off effect based on predefined thresholds of input variables were modified to have a sequential increase or decrease effect or a specific regression relation in order to increase the sensitivity to the variations of the input variables. This also resulted in a more detailed calibration process for each parameter.
For the calibration of the EPIRICE Daily Risk Model, rice blast incidence data collected from the RDA test plots of Bonghwa and Namyangju, South Korea, in 2004 were used. These two datasets were selected because they had a relatively good fit with the disease progress curves simulated by the EPIRICE model in our previous study (Kim et al., 2015). The disease progress curve of the rice blast is typically bell-shaped, and usually starts to appear around the 20th day after transplanting, showing the highest severity between 30 and 10 days before water withdrawal from rice paddies, and then gradually decreases. This is explained by the fact that the diseased leaves age and eventually fall down over time, being replaced by new healthy leaves, so the proportion of the disease lesions in the total leaf area decreases. For the calibration study, we used the increased severity between each disease survey point as a newly infected proportion during each disease survey period. Then, the increased severity was correlated with the weather conditions during the same period. As the severity decreases after reaching its highest peak, the increased severity thereafter becomes zero. However, we utilized all increased severity values, including the zero values, for the model calibration.
The detailed process for calibration of the EPIRICE Daily Risk Model was as follows: (1) from the disease incidence data of rice blast collected from the RDA test plots at 10-day intervals, the increased severity values between each disease survey point of the actual incidence data were calculated and displayed over days after transplanting; (2) the EPIRICE Daily Risk Model was run using historical weather data to generate daily risk scores; (3) the daily risk scores were divided into the periods during which the abovementioned increased severity had been calculated, and the daily risk scores in each period were summed; (4) Pearson correlation analyses with the increased severity values and the model-simulated sums of daily risk scores were performed for the Bonghwa and Namyangju datasets from 2004; (5) finally, the model calibration was repeated by micro-adjusting relevant parameters until the correlation coefficients (CC) for both sites reached 0.92 or higher (i.e., to show a significant correlation at 99% confidence level when 6 samples are used). Although the small number of samples did not mean much to the correlation analysis, it was regarded as a set goal to show that the model parameters had been corrected at the level that best reflects the actual disease incidences.
Validating the EPIRICE Daily Risk Model against ground-truth disease incidences.
To predict the potential blast risk for the coming season using SCFs and the EPIRICE Daily Risk Model, it is necessary to infer the actual disease severity level in rice paddies from the model output. In other words, the daily risk scores from the model need to be converted into realistic and measurable disease values in the field. As a first step to make the conversion function, we had to determine what information to use in the actual disease incidence data, which was then followed by the process of linking the information to the outputs of the model based on statistical correlation.
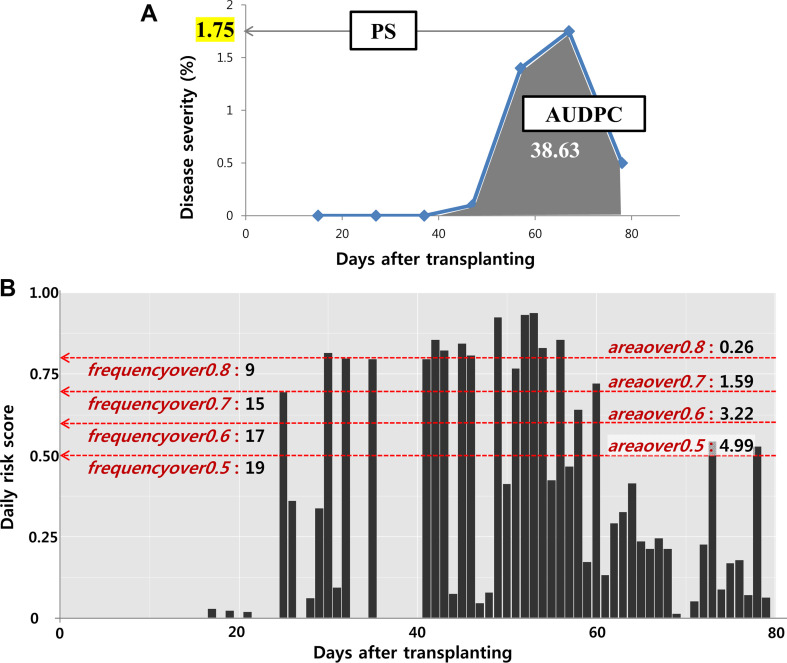
First, we used the disease incidence data of the RDA test plots from 2002 to 2010, which we had utilized for the parameterization and validation of the EPIRICE model in our previous study (Kim et al., 2015). For the correlation analysis, we selected the incidence data with the maximum peak severity (PS) of more than 0.5% because the lowest severity value that can be recorded by the surveyor is of 0.5%. In addition, only the datasets with at least one intermediate severity value recorded between the first severity value greater than zero and the maximum PS value were selected. Finally, the datasets with a typical rice blast progress curve shape (bell-shape) were selected. As a result of these quality controls (QC), a total of 13 rice blast incidence datasets were selected and used in the study. Then, we had to decide which information from the selected incidence datasets to use for the correlation analysis. The incidence datasets contained multiple survey points over time, with severity values measured at 10-day intervals, with which we drew a disease progress curve over time. In general, to determine the overall severity of a disease during a season, we use either the area under the disease progress curve (AUDPC) or the PS. Fig. 2A shows an example of the rice blast progress curve of one of the 13 incidence datasets, namely the Bonghwa dataset from 2004, for which we calculated an AUDPC value of 38.63 and a PS value of 1.75. In the same way, the AUDPC and PS values for the remaining 12 incidence datasets were calculated for the correlation analysis.
Fig. 2B shows the daily risk scores simulated by the EPIRICE Daily Risk Model using the observed weather and rice cultivar input data of the Bonghwa dataset from 2004. In order to correlate the simulated risk scores with the observed incidence data, we processed the daily risk scores into secondary indices that are compatible with the AUDPC or PS from the observed incidence data. First, we set multiple thresholds for the daily risk score, which were increased from 0.5 to 0.8 in 0.1 increments. This range was selected because when the threshold was below 0.5 or above 0.8, no values for statistical analysis were generated. By increasing the threshold value from 0.5 to 0.8, two indices, namely frequencyover and areaover, were calculated for each threshold. The frequencyover index is the frequency at which the daily risk scores exceed the threshold, and areaover is the sum of all the daily risk scores that are over the threshold. Each combination (or crossing pair) was named by combining the index (either frequencyover or areaover) and the threshold (0.5, 0.6, 0.7, or 0.8). For example, the frequencyover0.5 is the frequency of the daily risk scores over the threshold of 0.5. Based on the Pearson correlation analysis using the multiple crossing pairs with the AUDPC or PS values and the frequencyover or areaover indices, we identified the best combination of the actual severity level in rice paddies and the daily risk scores from the model. Using the best combination pair, a linear regression analysis was conducted to derive a regression equation, which was later used as a conversion function of the EPIRICE Daily Risk Model. The final conversion function was evaluated using the root-mean-square error (RMSE) between the observed PS values and the ones simulated by the conversion function of the model. The RMSE is the most commonly used measure of the discrepancies between observed values and model simulations because it represents the precision of the model when model simulations are compared to the observed values, and is defined as follows:
In Eq. (2), xi is the simulated PS value, xo is the observed PS value, and n is the number of samples, with a value of 13 in the present evaluation.
The input values for running the EPIRICE Daily Risk Model include daily weather data, namely air temperature, precipitation, and relative humidity, as well as the transplanting date and rice cultivar. Considering that the RDA test plot data cover the period from 2002 to 2010, the model was run using the available 1-km grid weather data from the South Korean Meteorological Administration (KMA). The KMA provides 1-km scale, gridded weather data of the South Korean region for the period of 2000-2010 (http://www.kma.go.kr), which have been generated from historic weather data collected from 76 automatic synoptic observation systems (ASOS) and 462 automatic weather stations (AWS) throughout South Korea, using the parameter-elevation regression on independent slopes model (PRISM) (Kim et al., 2012). Since the PRISM weather data included only temperature and precipitation data, we derived the relative humidity data from the observations of the closest ASOS and AWS. As for the information on the rice cultivars and transplanting dates, we used the data presented by Kim et al. (2015).
For the final validation of the EPIRICE Daily Risk Model with the new conversion function, we had to secure a set of ground-truth disease incidence data other than the RDA test plot data that were used for the conversion function development. The RDA had run a rice blast experiment plot with double-volume nitrogen fertilizer application in Icheon, Gyeonggi-do, South Korea, from 1974 to 2000, where the number of leaf blast lesions was counted on a weekly basis by the RDA plant pathologists. Notably, this is one of the experiments with the most accurate, multi-year rice blast data in South Korea. We calculated the AUDPC and PS values directly from the lesion numbers, which did not need to be converted into severity as the purpose of this validation test was to analyze the correlation between the observed and simulated data. In addition, the weather input data for model simulation were collected from the Icheon ASOS station (ID 203) of the KMA, which is located less than 5 km from the Icheon rice blast experiment plot. Finally, we used Jinheung as a cultivar with moderately susceptible resistance and May 25 as the transplanting date to run the EPIRICE Daily Risk Model.
Sensitivity analysis of the EPIRICE Daily Risk Model.
To determine how sensitive the EPIRICE Daily Risk Model is to individual weather conditions, we performed sensitivity tests on all weather input variables. Daily temperature, precipitation, and relative humidity data corresponding to the disease incidence survey conducted in Bonghwa in 2004 were used as reference data. The model was simulated for 80 days from the transplanting date, May 25, to August 12. The simulated daily risk score was converted to the areaover0.6 value with the conversion function developed in our study. The sensitivity analysis examined the effects of each weather variable on the areaover0.6 value from the EPIRICE Daily Risk Model. The range of each weather variable for the sensitivity analysis was determined based on the past 30 years (i.e., 1976-2005) data of the KMA ASOS station (ID 271) in Bonghwa. The lowest and highest values recorded over this 30-year period were obtained for each weather variable (i.e., mean temperature: ~19.8-24.3┬░C; mean relative humidity: ~64.5-84.3%; cumulative precipitation: ~218.5-917.5 mm), and the ranges used for the analysis were determined between 95% of the lowest value and 105% of the highest value.
Results
Structure of the EPIRICE Daily Risk Model.
The EPIRICE Daily Risk Model converts daily weather data into daily rice blast risk scores based on four modules, namely temperature, leaf wetness, plant age, and cultivar susceptibility (Fig. 3A). By separating the original temperature function from the infection algorithm of the EPIRICE model, we developed a temperature module that determines the relative disease risk according to the average daily temperature (Fig. 3B). It shows the highest relative risk by temperature, namely the T effect, at approximately 21┬░C. Under temperature conditions below 9┬░C or above 38┬░C, where the T effect becomes 0, the daily infection rate also becomes 0. This principle applies to all four modules of the EPIRICE Daily Risk Model, which means that the daily infection rate becomes 0 whenever any of the effect values becomes 0. Notably, this mimics the actual conditions of disease development. That is, when one essential condition is not suitable for the occurrence of the disease, the condition becomes a limiting factor and the disease does not usually occur.
In the EPIRICE model, the algorithm for the effect of relative humidity and precipitation on the infection rate is quite simplified. When the maximum relative humidity of the day exceeds 90% or the daily precipitation exceeds 5 mm, the effect of the relative humidity and precipitation becomes 1; otherwise, its value is of 0. In the EPIRICE Daily Risk Model, this was improved to a more detailed function in order to estimate leaf wetness (Fig. 3D). The daily precipitation and relative humidity were used to calculate the relative disease risk in relation to leaf wetness, namely the leaf wetness (LW) effect. First, we specified the range of daily precipitation in terms of their relative contribution to the daily infection rate. Precipitations of 1 to 50 mm, which in general help the infection by maintaining leaf wetness, cause the highest relative disease risk. As for precipitations between 50 and 100 mm, which remove the fungal inocula of rice blast from the air, cause the LW effect to drop by half. Finally, precipitations greater than 100 mm suppress disease occurrence by washing down all blast inocula that are in the air or on the plant surface, so the LW effect becomes 0. Even with less than 1 mm of daily precipitation, however, the infection can take place under a leaf wetness condition influenced by high relative humidity in the air. Therefore, the leaf wetness duration was estimated based on the daily diurnal variation of relative humidity using the latitude, date, average relative humidity, and the minimum and maximum temperatures at the site (Hijmans, 2016; Wichink Kruit et al., 2004). The LW effect was then determined based on the estimated leaf wetness duration (El Refaei, 1977; Singh and Mohan, 2016), as shown in Fig. 3D.
We had adopted a simple algorithm for adult plant tolerance to rice blast in the EPIRICE model and, hence, developed a plant age module in the EPIRICE Daily Risk Model (Fig. 3C). The plant age module exhibits relative disease risk by plant age, namely the PA effect, by establishing an empirical relation that changes the susceptibility of rice plants to rice blast according to plant age (i.e., the number of days after transplanting). To simplify this compared to other modules, the PA effect is maintained at a value of 1 up to 40 days after transplanting, after which it gradually drops to 0 until 70 days after transplanting. Lastly, the EPIRICE model has a host parameter with five general degrees of host resistance to blast disease: susceptible, moderately susceptible, neutral, moderately resistant, and resistant. Based on this, we developed the cultivar susceptibility module of the EPIRICE Daily Risk Model (Fig. 3E). The cultivar susceptibility module shows the relative disease risk by rice cultivar, namely the CS effect. As it never falls down to 0, unlike other modules, the CS effect acts as a base infection rate that is constantly adjusted by other effects on a daily basis.
Calibration of the EPIRICE Daily Risk Model.
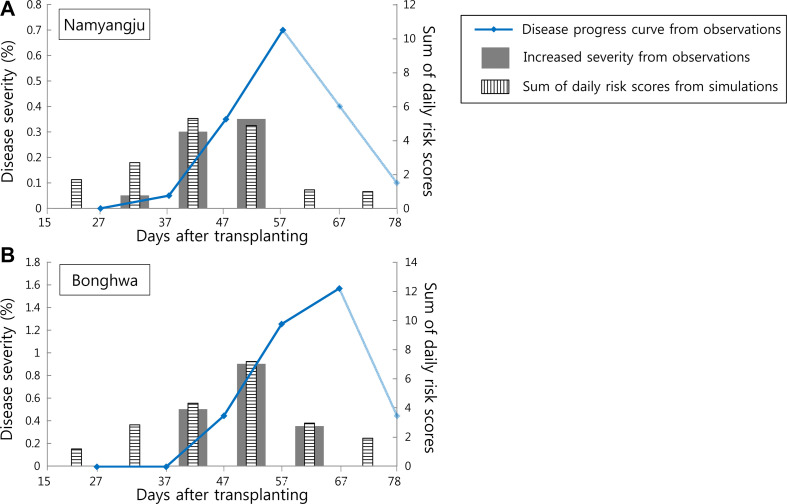
After multiple rounds of calibrating the parameters of the EPIRICE Daily Risk Model, we successfully reached final Pearson CC of more than 0.92 for both sites. Fig. 4 shows the resulting daily risk scores from the calibrated EPIRICE Daily Risk Model, driven by the final parameters, and the corresponding RDA test plot data of both sites. The temporal changes in the increased severity at each site and the simulated sum of daily risk scores showed a good graphical agreement. The final CCs were of 0.962 and 0.948 for Namyangju and Bonghwa, respectively. Therefore, the calibrated EPIRICE Daily Risk Model was considered to produce a simulation of the temporal development of rice blast severity that is representative of the corresponding development in the field. The resulting parameters of the calibrated model are shown in the modules B-E of Fig. 3.
Validation of the EPIRICE Daily Risk Model.
The final results of the Pearson correlation analysis (Table 1) indicate that the combination of the PS value and areaover0.6 index showed the highest CC, with a value of 0.949 at the 95% confidence level. Although some combinations had similar CC values, such as 0.935 for the PS with areaover0.5, 0.917 for the PS with areaover0.7, and 0.914 for the PS with frequencyover0.8, we finally selected the combination of the PS and areaover0.6 for the next analysis. Notably, the CCs were always higher with the PS than with the AUDPC values in all crossing pairs. The linear regression analysis using the 13 pairs of the PS and the areaover0.6 data resulted in a regression equation of y = 0.6467 ├Ś x + 0.6128 (y, PS; x, areaover0.6) and an R2 of 0.9. This equation was then used as a conversion function in the EPIRICE Daily Risk Model to convert the model outputs into the PS of blast disease during a season. The RMSE between the observed PS values from the 13 blast incidence datasets from the RDA test plots and the simulated PS values from the conversion function of the model was 0.47%, which is less than the lowest severity value of 0.50% that can be identified by a surveyor, and was thus considered as a negligible error from using the conversion function.
As for the validation of the calibrated EPIRICE Daily Risk Model and the new conversion function, the Pearson CC between the observed peak number of leaf blast lesions and the simulated areaover0.6 indices from the model was 0.7 at the 95% confidence level. Based on this result, we concluded that the EPIRICE Daily Risk Model with the conversion function performs reasonably well against the observed rice blast incidence data collected at multiple sites in South Korea, which indicates that the EPIRICE Daily Risk Model can later be used to generate seasonal blast risk information for the seasonal disease early warning service.
Sensitivity of the EPIRICE Daily Risk Model to weather input variables.
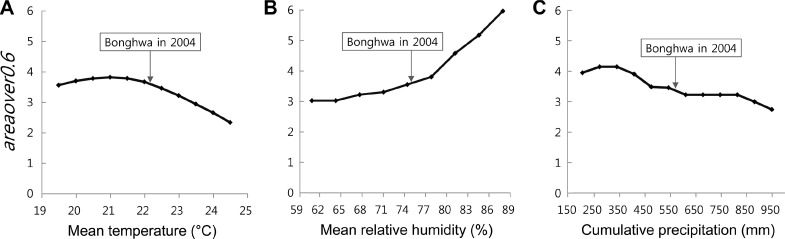
A sensitivity analysis characterizes the response of model outputs to input variation. Based on the sensitivity test, the responses of the EPIRICE Daily Risk Model to weather input variables were determined. The reference weather values of Bonghwa from 2004 (Fig. 5) showed that the reference conditions were approximately median within the annual variation of each weather variable over the past 30 years. The weather variable that showed the most sensitive response in the EPIRICE Daily Risk Model was the relative humidity, for which the difference between the highest and lowest areaover0.6 values was more than 3. A temperature shift from 19 to 25┬░C resulted in the areaover0.6 values changing from 2.3 to 3.9, while the increase in precipitation from ~200 to 950 mm changed the areaover0.6 values from 2.7 to 4.2. In the case of the relative humidity, the areaover0.6 values slowly increased in relation to the relative humidity range of approximately 60 to 80%, and then they increased significantly as the relative humidity exceeded 80%. This is presumably explained by the fact that the LW module has been parameterized to have a significant impact on the daily infection rate within a specific range of relative humidity, which is generally relevant to leaf wetness formation in the field.
Discussion
In this study, we developed the EPIRICE Daily Risk Model, which can immediately utilize downscaled SCFs, in an effort to overcome the limitations of the previous EPIRICE model. The original purpose of the EPIRICE model was to predict the potential disease risk by using the AUDPC, which implied that the model output could be obtained only when weather data for the whole season was available. Therefore, it was difficult to produce disease risk predictions for only a certain period within the season, which is required when using SCFs for a certain period only. Furthermore, there was a structural complexity in the EPIRICE when considering the relationship between the growth of crops and disease development. In this study, the following modifications were made not only to increase the overall simplicity of the model, but also to simulate daily infection risks, which primarily depend on daily weather conditions. Firstly, the central crop growth and senescence modules, as well as the susceptible-exposed-infectiousremoved infection module, of the EPIRICE model have been completely removed. In the remaining infection rate element of the EPIRICE model, the number of infectious sites and the disease aggregation function (Ca), which is the rate of spreading from the infected area to the surrounding healthy plants, have been removed, only leaving Rc, the basic infection rate. For detailed descriptions of the modules and elements from the EPIRICE model that we have removed, readers are referred to Savary et al. (2012) and Kim et al. (2015).
The daily risk scores from the EPIRICE Daily Risk Model refer to the degree of risk that pathogens will infect healthy rice if there are sufficient inocula present. For rice that has already been infected, the score corresponds to how quickly pathogens colonize the infected plant site, or how quickly a new source of infection (e.g., fungal conidia) spreads from the infected site to nearby plants. Therefore, this risk information can be utilized for diseases that require preventive control. For example, proper management actions can be applied in advance, before the risk exceeds a certain level where prevention through proactive control is no longer effective (Berger, 1977). In order to apply this principle, it was necessary to find a link between the daily risk scores produced by the EPIRICE Daily Risk Model and the level of rice blast incidence in the field. Accordingly, the conversion function that we developed in the study converts the model outputs into more epidemiologically relevant information, namely the PS of rice blast in the field. The PS generated by the model can be directly used by decision-makers to decide whether to apply chemical sprays, considering the economic threshold (ET) of rice blast. Yeh et al. (2008) reported that the ET for rice yield is 2.7% of the PS, which corresponds to the time when the application of fungicide is necessary. Considering the calendar-based fungicide applications for collaborative disease controls in South Korea, some of the applications might be unnecessary under very low disease pressure. Thus, linking the PS estimation from the EPIRICE Daily Risk Model with the ET information for each crop-disease combination will enable the calculation of a cost-benefit ratio and increase disease management efficiency, by assisting agricultural stakeholders to make science-based, informed decisions.
The conversion function of the EPIRICE Daily Risk Model is based on our understanding of the epidemiological perspective on diseases in relation to conducive weather conditions. Simply using the cumulative values of all the daily risk scores or using the average value of the risk scores may not be an appropriate approach because there are many occasions where the infection does not progress under a certain threshold (De Wolf et al., 2003; Madden and Ellis, 1988; Xu, 2003). In order for a pathogen to infect a plant or spread to surrounding healthy plants, there must be an inducing environment above a certain level (i.e., intensity and duration). For example, even if the temperature is exactly within the optimal range, the disease does not develop unless the relative humidity, precipitation, or other non-environmental factors become suitable for the infection. In fact, our correlation analysis for the conversion function revealed that our assumption that only daily risk scores above a certain level conduce the disease development or spread was reasonable.
In this study, despite the fairly high CCs from the conversion and validation experiments, there is still a limitation in that the number of samples was relatively small and the 13 disease incidence datasets do not sufficiently represent the entire rice paddies of South Korea. Although more ground-truth disease incidence data for model validation are needed, it is very difficult to obtain additional data in South Korea, as shown in the RDA test plots, where the occurrence of rice blast since 2010 has been negligible (Rural Development Administration, 2015). Furthermore, there are many factors influencing the occurrence of rice blast in addition to the weather and cultivar type, such as the availability of inoculum, agronomic practices, and human factors (Heong et al., 1995; Long et al., 2000). Another limitation is the quality of the disease incidence data, primarily related to the surveyor-dependent quality variation, although our QC processes had already ruled out some low-quality field data. As Rykiel (1996) stated, a model cannot be expected to generate results that are more accurate and precise than data from the actual system. In other words, the testability of a model is defined by the accuracy and precision of the disease incidence data used for model development. Therefore, continuous, high-quality, and standardized data collection through field trials is required, under the condition that all non-environmental factors are controlled.
Based on the sensitivity analysis, the responses of the EPIRICE Daily Risk Model to weather input variables were examined. The sensitivity analysis of each weather variable indicates that the model algorithm is sensitive enough to respond not only to the overall effects of weather conditions but also to individual weather conditions, which is in line with the blast epidemiological mechanisms that have been revealed by many studies (Agrios, 2005; Calvero et al., 1996; Kim and Kim, 1993). This also illustrates that we carefully selected the variables and their representative ranges for the analysis (Vonk Noordegraaf et al., 2003). Among all weather variables, the relative humidity resulted in the most sensitive (i.e., greatest) changes in the model outputs, which is consistent with the fact that leaf wetness is one of the most critical factors for the successful germination of and penetration by the blast fungus (Greer and Webster, 2001; Katsantonis et al., 2017). In the case of cumulative precipitation, the sensitivity analysis was performed by changing the intensity of precipitation from the Bonghwa dataset while maintaining the precipitation frequency. Although our initial assumption was that precipitation amount variations would not significantly affect the sensitivity of the model, the areaover0.6 value of the model responded fairly sensitively and ranged from 2.7 to 4.2. A possible explanation is that the leaf wetness module of the EPIRICE Daily Risk Model was parameterized to generate differential LW effect values according to the intensity of precipitation.
Assuming that the available SCFs provide prediction information up to 3-month lead time (Easterling and Mjelde, 1987), all necessary inputs have to be entered into the model at the end of May to generate a seasonal rice blast risk index (in the form of the PS) for June-August. Based on the seasonal blast risk information, it will be possible for users to prepare various strategic or tactical countermeasures in advance. As the SCFs will be spatially downscaled into higher resolution grids distributed throughout South Korea, it is also possible for local and central government entities to make national-level policy decisions. In this way, the seasonal disease early warning service can be complementary to the NCPMS of the RDA. In addition, timely disease predictions will not only induce control at the right time, but also result in the reduction of the total management cost by avoiding unnecessary sprays, which will also decrease the side effects of the excessive use of chemical sprays on the environment and human health, among others. This is an unrealized but potential value of seasonal disease early warning services (Roberts et al., 2006). Overall, we developed and validated the EPIRICE Daily Risk Model to provide a baseline for follow-up efforts. If the value of SCFs is revealed and the necessity of seasonal disease predictions increases in the future, many researchers in related fields will be able to participate through joint research and collaboration. As a result, if such a common effort is initiated, there may be sufficient pest and disease models, capable of operationalizing seasonal disease early warning services by using SCFs, in the near future.





 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print






